RandomForest with scikit-learn#
We want to explore how to use scikit-learn for RandomForest. This includes preparing the data, training the model, and evaluating and visualizing the results.
We will use the following modules for this:
sklearn.datasets: Tools for using common datasets for ML or for generating synthetic data
sklearn.model_selection: Tools for data splitting, cross-validation, and parameter tuning
sklearn.tree and sklearn.ensemble: Collection of tree- and ensemble-based models for regression and classification
sklearn.metrics: Collection of various metrics for model evaluation
In general, also refer to the comprehensive User Guide on decision trees and User Guide on ensemble models from scikit-learn.
# Just in case we need help
# Import bia-bob as a helpful Python & Medical AI expert
import os
from bia_bob import bob
bob.initialize(
endpoint=os.getenv('ENDPOINT_URL'),
model="vllm-llama-4-scout-17b-16e-instruct",
system_prompt=os.getenv('SYSTEM_PROMPT_MEDICAL_AI')
)
%bob Who are you ? Just 1 sentence!
I’m an expert in medical data science and a skilled Python programmer and data analyst with extensive experience working with various medical datasets and applying data analysis, machine learning, and deep learning techniques.
RandomForest for Regression#
Data preparation#
Now we want to use a RandomForest to solve the regression problem for the scikit-learn diabetes dataset.
Let’s load the data again:
from sklearn.datasets import load_diabetes
diabetes = load_diabetes()
# Let's get our X,y first
X = diabetes.data
y = diabetes.target
# And here we also have proper labels for our features
feature_labels = diabetes.feature_names
print("Type X:", type(X))
print("Shape X:", X.shape)
print("First X:", X[0])
print("Labels X:", feature_labels)
Type X: <class 'numpy.ndarray'>
Shape X: (442, 10)
First X: [ 0.03807591 0.05068012 0.06169621 0.02187239 -0.0442235 -0.03482076
-0.04340085 -0.00259226 0.01990749 -0.01764613]
Labels X: ['age', 'sex', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']
print("Type y:", type(y))
print("Shape y:", y.shape)
print("First y:", y[0])
Type y: <class 'numpy.ndarray'>
Shape y: (442,)
First y: 151.0
Train-test split#
Now we split the data for training and model testing / evaluation in order to verify the generalization of the trained model on unknown data. For this purpose, we can use train_test_split.
from sklearn.model_selection import train_test_split
# Split: 80% training, 20% test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
print("training data:", X_train.shape, "training labels:", y_train.shape)
print("test data:", X_test.shape, "test labels:", y_test.shape)
training data: (353, 10) training labels: (353,)
test data: (89, 10) test labels: (89,)
Select and train a model#
We import the model
RandomForestRegressorand initialize the corresponding Python object using()while also setting specific model parametersModel training is started using the method
.fit()We pass the training data to this method, divided into features
X_trainand targety_trainThe model parameters are now adjusted to predict the target as accurately as possible based on the associated feature
from sklearn.ensemble import RandomForestRegressor
# Initialize model with model parameters
# - n_estimators: number of decision trees in the forest
# - max_depth: maximum depth of a tree
model = RandomForestRegressor(
n_estimators=5,
max_depth=None
)
# Supervised training - “Fitting” the model to the training data with known labels
model.fit(X_train, y_train)
RandomForestRegressor(n_estimators=5)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Prediction and evaluation#
We now want to check how well the model can predict the target for previously unknown features.
The prediction is performed using the method
.predict()– almost all models and algorithms in scikit-learn have this method.We pass the features of the test data to this method and receive the corresponding predictions of the target.
# Prediction on unseen test data
y_pred = model.predict(X_test)
We can now compare the predictions with the known target values of the test data and derive metrics for evaluating the model quality.
A metric provided by RandomForestRegressor itself is the so-called
r2score (coefficient of determination of a regression), which evaluates the overall goodness of fitOther suitable regression metrics are described, for example, in the User Guide - Regression Metrics
We also choose the Mean Absolute Error (MAE)
In general, it makes sense to calculate several metrics in order to get a better impression of the model quality
from sklearn.metrics import mean_absolute_error
# Metrics for determining model quality
# r^2 score, between 0.0 and 1.0, higher is better
r2_score = model.score(X_test, y_test)
print(f"r^2 score on test data: {r2_score:.3f}")
# Calculate mean absolute error (MAE), the best is 0.0
mse = mean_absolute_error(y_test, y_pred)
print(f"Mean Absolute Error (MAE) on test data: {mse:.3f}")
r^2 score on test data: 0.379
Mean Absolute Error (MAE) on test data: 47.007
Parameter tuning#
At the moment, we have simply estimated the values for the model parameters n_estimators and max_depth and obtained an evaluation result for this combination. But how can we be sure that this is actually the best choice, or whether we can still improve the model quality?
# We can check all available parameters again
model.get_params()
{'bootstrap': True,
'ccp_alpha': 0.0,
'criterion': 'squared_error',
'max_depth': None,
'max_features': 1.0,
'max_leaf_nodes': None,
'max_samples': None,
'min_impurity_decrease': 0.0,
'min_samples_leaf': 1,
'min_samples_split': 2,
'min_weight_fraction_leaf': 0.0,
'monotonic_cst': None,
'n_estimators': 5,
'n_jobs': None,
'oob_score': False,
'random_state': None,
'verbose': 0,
'warm_start': False}
Since trial and error would be very time-consuming with a large number of available parameters and combinations, we should use an automated approach that employs scikit-learn hyper-parameter tuning methods.
We want to apply a basic grid search over specified parameter values using sklearn.model_selection.GridSearchCV
from sklearn.model_selection import GridSearchCV
# Define the parameter grid we want to search
param_grid = {
'n_estimators': [1, 25, 50, 200],
'max_depth': [None, 5, 25, 50],
}
# Initialize GridSearchCV with the following parameters
# - estimator: the model we want to tune
# - param_grid: the parameter grid to search from
# - scoring: score used to determine the model quality for given parameters
# - cv: cross-validation data splitting strategy (n-fold)
grid_search = GridSearchCV(
estimator=RandomForestRegressor(random_state=42),
param_grid=param_grid,
scoring='r2',
cv=5
)
# Perform grid search, may take a while
grid_search.fit(X_train, y_train)
GridSearchCV(cv=5, estimator=RandomForestRegressor(random_state=42),
param_grid={'max_depth': [None, 5, 25, 50],
'n_estimators': [1, 25, 50, 200]},
scoring='r2')In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
RandomForestRegressor(max_depth=5, n_estimators=200, random_state=42)
Parameters
# Print best parameter combination with the best score
print("Best parameters for training data:", grid_search.best_params_)
print("Best score for training data:", grid_search.best_score_)
# Get the best model with the best parameter choice
best_model = grid_search.best_estimator_
Best parameters for training data: {'max_depth': 5, 'n_estimators': 200}
Best score for training data: 0.40719110568079026
As we can see, setting max_depth=5 and n_estimators=200 delivered the best results for the training data.
Let’s evaluate the predictions of this best model with our test data.
# Prediction on unseen test data
y_pred_best = best_model.predict(X_test)
# r^2 score, between 0.0 and 1.0, higher is better
r2_score = best_model.score(X_test, y_test)
print(f"r^2 score on test data: {r2_score:.3f}")
# Calculate mean absolute error (MAE), the best is 0.0
mse = mean_absolute_error(y_test, y_pred_best)
print(f"Mean Absolute Error (MAE) on test data: {mse:.3f}")
r^2 score on test data: 0.460
Mean Absolute Error (MAE) on test data: 43.274
Explainability#
Tree insights#
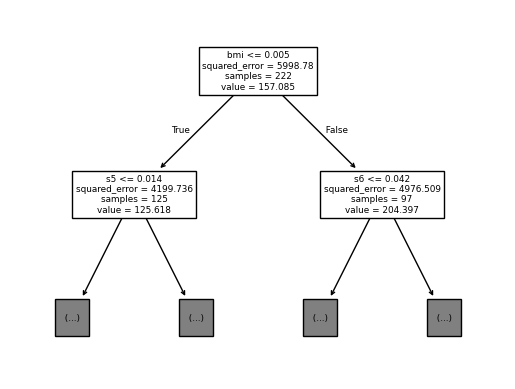
Now that we have trees built inside our RandomForest, we can examine separate trees by plotting them.
from sklearn import tree
# Plot the first tree from our RandomForest with limited depth
tree.plot_tree(best_model.estimators_[0],
max_depth=1,
feature_names = feature_labels)
[Text(0.5, 0.8333333333333334, 'bmi <= 0.005\nsquared_error = 5998.78\nsamples = 222\nvalue = 157.085'),
Text(0.25, 0.5, 's5 <= 0.014\nsquared_error = 4199.736\nsamples = 125\nvalue = 125.618'),
Text(0.375, 0.6666666666666667, 'True '),
Text(0.125, 0.16666666666666666, '\n (...) \n'),
Text(0.375, 0.16666666666666666, '\n (...) \n'),
Text(0.75, 0.5, 's6 <= 0.042\nsquared_error = 4976.509\nsamples = 97\nvalue = 204.397'),
Text(0.625, 0.6666666666666667, ' False'),
Text(0.625, 0.16666666666666666, '\n (...) \n'),
Text(0.875, 0.16666666666666666, '\n (...) \n')]
Feature importance#
Now that we have an example with more than one feature, we can also evaluate the significance / importance of the features for the regression task. This helps to understand which features contributed to the prediction and to what extent. This information can then be used for a more refind feature selection.
For this, some models in scikit-learn provide the attribute feature_importances_ (impurity-based, good for low cardinality features). In addition, the more sophisticated method sklearn.inspection.permutation_importance is available.
import numpy as np
import pandas as pd
# Extract feature importances from the best model, store sorted in a pandas Series
importances = pd.Series(best_model.feature_importances_, index=feature_labels).sort_values(ascending=False)
# Get the standard deviations of importances across trees as a measure of the variability
importances_std = np.std([tree.feature_importances_ for tree in best_model.estimators_], axis=0)
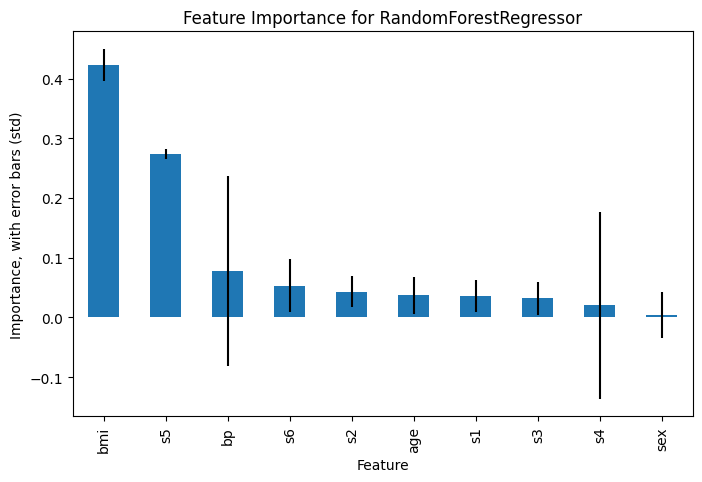
Now we can plot the feature importances, including the standard deviations as error bars
import matplotlib.pyplot as plt
plt.figure(figsize=(8, 5))
importances.plot(kind='bar', yerr=importances_std)
plt.xlabel('Feature')
plt.ylabel('Importance, with error bars (std)')
plt.title('Feature Importance for RandomForestRegressor')
plt.show()
Exercise: Classification with RandomForest#
Now it’s your turn to apply a RandomForest to a classification problem, using the scikit-learn breast cancer diagnostic dataset.
This data contains features computed from 569 digitized images of a breast mass, which shall be used to predict two classes: WDBC-Malignant, or WDBC-Benign.
For example, you can use the following objects and methods:
RandomForestClassifieras modelGridSearchCVorRandomizedSearchCVfor parameter optimizationClassification Metrics to evaluate the model quality
permutation_importancefor explainability
If you get stuck, remember that our assistant bia-bob is available and very happy to help you.
Data preparation#
from sklearn.datasets import load_breast_cancer
breast_cancer = load_breast_cancer()
dir(breast_cancer)
['DESCR',
'data',
'data_module',
'feature_names',
'filename',
'frame',
'target',
'target_names']
# ToDo: understand the data better
Train-test split#
# ToDo: create data for training and test
Select and train a model#
# ToDo: initialize and train a classification model with parameter optimization
Prediction and evaluation#
# ToDo: evaluate the model quality
