CNNs for radiology images#
In this notebook we will apply our new CNN knowledge to classify medical images. For this, we use a chest X-ray dataset from MedMNIST. Here, we want to classify patients into healthy and pneumonia cases. It was originally derived from a larger pediatric chest X-ray dataset.
Goals:
load X-ray data
build and train CNN for pneumonia classification
evaluate the model
have a first look at explainability
from google.colab import ai
response = ai.generate_text("Tell me what AI you are and can you help me with coding problems?")
print(response)
Hello!
I am a large language model, trained by Google. My purpose is to assist with a wide range of tasks by processing information and generating human-like text based on the vast amount of data I've been trained on. I don't have personal experiences, consciousness, or feelings; I'm an AI designed to communicate and provide information.
And yes, absolutely! **I can definitely help you with coding problems.**
Here's how I can assist you:
* **Explaining Concepts:** I can break down complex programming concepts, algorithms, data structures, and design patterns.
* **Writing Code Snippets:** I can generate code snippets, functions, or even small programs in various programming languages (Python, JavaScript, Java, C++, Ruby, Go, etc.) based on your requirements.
* **Debugging Assistance:** If you have an error, you can share your code and the error message, and I can help you identify the potential cause and suggest solutions.
* **Code Review and Improvement:** I can look at your code and suggest ways to make it more efficient, readable, or adhere to best practices.
* **Syntax and Language Help:** If you're stuck on how to perform a specific action in a language, I can provide the correct syntax and examples.
* **Algorithm Design:** I can help you brainstorm different approaches to solve a problem and explain the pros and cons of various algorithms.
* **Learning Resources:** I can point you towards documentation, tutorials, or other learning materials for specific languages or technologies.
* **Framework and Library Usage:** I can explain how to use popular frameworks and libraries and provide examples.
**To get the best help, please provide:**
* **The programming language** you're using.
* **Your code snippet** (if applicable).
* **The error message** (if applicable).
* **What you're trying to achieve** or what the problem is.
* **What you've tried so far.**
So, feel free to ask away! Just tell me what you're working on, and I'll do my best to assist you.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader, TensorDataset
from torchvision import transforms
import torch.nn.functional as F
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import roc_auc_score, roc_curve, confusion_matrix, ConfusionMatrixDisplay
from sklearn.metrics import classification_report, accuracy_score, precision_score, recall_score, f1_score
1) Load MedMNIST datset#
We use the medmnist package, which provides programmatic access to MedMNIST cohorts.
!pip install medmnist # should take around 2-3 min
import medmnist
from medmnist import INFO
Requirement already satisfied: medmnist in /usr/local/lib/python3.12/dist-packages (3.0.2)
Requirement already satisfied: numpy in /usr/local/lib/python3.12/dist-packages (from medmnist) (2.0.2)
Requirement already satisfied: pandas in /usr/local/lib/python3.12/dist-packages (from medmnist) (2.2.2)
Requirement already satisfied: scikit-learn in /usr/local/lib/python3.12/dist-packages (from medmnist) (1.6.1)
Requirement already satisfied: scikit-image in /usr/local/lib/python3.12/dist-packages (from medmnist) (0.25.2)
Requirement already satisfied: tqdm in /usr/local/lib/python3.12/dist-packages (from medmnist) (4.67.3)
Requirement already satisfied: Pillow in /usr/local/lib/python3.12/dist-packages (from medmnist) (11.3.0)
Requirement already satisfied: fire in /usr/local/lib/python3.12/dist-packages (from medmnist) (0.7.1)
Requirement already satisfied: torch in /usr/local/lib/python3.12/dist-packages (from medmnist) (2.10.0+cu128)
Requirement already satisfied: torchvision in /usr/local/lib/python3.12/dist-packages (from medmnist) (0.25.0+cu128)
Requirement already satisfied: termcolor in /usr/local/lib/python3.12/dist-packages (from fire->medmnist) (3.3.0)
Requirement already satisfied: python-dateutil>=2.8.2 in /usr/local/lib/python3.12/dist-packages (from pandas->medmnist) (2.9.0.post0)
Requirement already satisfied: pytz>=2020.1 in /usr/local/lib/python3.12/dist-packages (from pandas->medmnist) (2025.2)
Requirement already satisfied: tzdata>=2022.7 in /usr/local/lib/python3.12/dist-packages (from pandas->medmnist) (2025.3)
Requirement already satisfied: scipy>=1.11.4 in /usr/local/lib/python3.12/dist-packages (from scikit-image->medmnist) (1.16.3)
Requirement already satisfied: networkx>=3.0 in /usr/local/lib/python3.12/dist-packages (from scikit-image->medmnist) (3.6.1)
Requirement already satisfied: imageio!=2.35.0,>=2.33 in /usr/local/lib/python3.12/dist-packages (from scikit-image->medmnist) (2.37.2)
Requirement already satisfied: tifffile>=2022.8.12 in /usr/local/lib/python3.12/dist-packages (from scikit-image->medmnist) (2026.2.24)
Requirement already satisfied: packaging>=21 in /usr/local/lib/python3.12/dist-packages (from scikit-image->medmnist) (26.0)
Requirement already satisfied: lazy-loader>=0.4 in /usr/local/lib/python3.12/dist-packages (from scikit-image->medmnist) (0.4)
Requirement already satisfied: joblib>=1.2.0 in /usr/local/lib/python3.12/dist-packages (from scikit-learn->medmnist) (1.5.3)
Requirement already satisfied: threadpoolctl>=3.1.0 in /usr/local/lib/python3.12/dist-packages (from scikit-learn->medmnist) (3.6.0)
Requirement already satisfied: filelock in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (3.24.3)
Requirement already satisfied: typing-extensions>=4.10.0 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (4.15.0)
Requirement already satisfied: setuptools in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (75.2.0)
Requirement already satisfied: sympy>=1.13.3 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (1.14.0)
Requirement already satisfied: jinja2 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (3.1.6)
Requirement already satisfied: fsspec>=0.8.5 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (2025.3.0)
Requirement already satisfied: cuda-bindings==12.9.4 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.9.4)
Requirement already satisfied: nvidia-cuda-nvrtc-cu12==12.8.93 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.8.93)
Requirement already satisfied: nvidia-cuda-runtime-cu12==12.8.90 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.8.90)
Requirement already satisfied: nvidia-cuda-cupti-cu12==12.8.90 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.8.90)
Requirement already satisfied: nvidia-cudnn-cu12==9.10.2.21 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (9.10.2.21)
Requirement already satisfied: nvidia-cublas-cu12==12.8.4.1 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.8.4.1)
Requirement already satisfied: nvidia-cufft-cu12==11.3.3.83 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (11.3.3.83)
Requirement already satisfied: nvidia-curand-cu12==10.3.9.90 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (10.3.9.90)
Requirement already satisfied: nvidia-cusolver-cu12==11.7.3.90 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (11.7.3.90)
Requirement already satisfied: nvidia-cusparse-cu12==12.5.8.93 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.5.8.93)
Requirement already satisfied: nvidia-cusparselt-cu12==0.7.1 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (0.7.1)
Requirement already satisfied: nvidia-nccl-cu12==2.27.5 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (2.27.5)
Requirement already satisfied: nvidia-nvshmem-cu12==3.4.5 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (3.4.5)
Requirement already satisfied: nvidia-nvtx-cu12==12.8.90 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.8.90)
Requirement already satisfied: nvidia-nvjitlink-cu12==12.8.93 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (12.8.93)
Requirement already satisfied: nvidia-cufile-cu12==1.13.1.3 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (1.13.1.3)
Requirement already satisfied: triton==3.6.0 in /usr/local/lib/python3.12/dist-packages (from torch->medmnist) (3.6.0)
Requirement already satisfied: cuda-pathfinder~=1.1 in /usr/local/lib/python3.12/dist-packages (from cuda-bindings==12.9.4->torch->medmnist) (1.4.0)
Requirement already satisfied: six>=1.5 in /usr/local/lib/python3.12/dist-packages (from python-dateutil>=2.8.2->pandas->medmnist) (1.17.0)
Requirement already satisfied: mpmath<1.4,>=1.1.0 in /usr/local/lib/python3.12/dist-packages (from sympy>=1.13.3->torch->medmnist) (1.3.0)
Requirement already satisfied: MarkupSafe>=2.0 in /usr/local/lib/python3.12/dist-packages (from jinja2->torch->medmnist) (3.0.3)
data = "pneumoniamnist"
info = INFO[data]
DataClass = getattr(medmnist, info["python_class"])
print(info["task"])
print(info["label"])
transform = transforms.Compose([
transforms.ToTensor()
])
binary-class
{'0': 'normal', '1': 'pneumonia'}
3) Train/validation/test split#
train_dataset = DataClass(split="train", transform=transform, download=True)
val_dataset = DataClass(split="val", transform=transform, download=True)
test_dataset = DataClass(split="test", transform=transform, download=True)
print(len(train_dataset), len(val_dataset), len(test_dataset))
4708 524 624
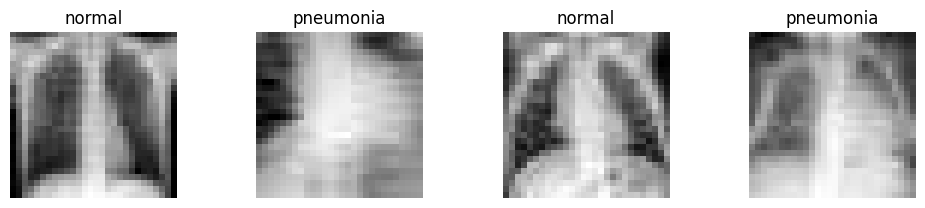
Lets have a look at some example images. As you can see the images are very pixelated (28x28 pixels). This is of course not the resulution used in clinical practice.
class_names = [info["label"][str(i)] for i in range(len(info["label"]))]
plt.figure(figsize=(10, 4))
for i in range(4):
img, label = train_dataset[i+5]
plt.subplot(2, 4, i + 1)
plt.imshow(img.squeeze(), cmap="gray")
plt.title(class_names[int(label)])
plt.axis("off")
plt.tight_layout()
plt.show()
/tmp/ipykernel_1796/2609169082.py:8: DeprecationWarning: Conversion of an array with ndim > 0 to a scalar is deprecated, and will error in future. Ensure you extract a single element from your array before performing this operation. (Deprecated NumPy 1.25.)
plt.title(class_names[int(label)])

BATCH_SIZE = 64
train_loader = DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True)
val_loader = DataLoader(val_dataset, batch_size=BATCH_SIZE, shuffle=False)
test_loader = DataLoader(test_dataset, batch_size=BATCH_SIZE, shuffle=False)
4) Define a CNN#
class CNN(nn.Module):
def __init__(self):
super().__init__()
self.features = nn.Sequential(
nn.Conv2d(1, 16, kernel_size=3, padding=1),
nn.ReLU(),
nn.MaxPool2d(2),
nn.Conv2d(16, 32, kernel_size=3, padding=1),
nn.ReLU(),
nn.MaxPool2d(2),
nn.Conv2d(32, 64, kernel_size=3, padding=1),
nn.ReLU(),
nn.AdaptiveAvgPool2d((1, 1))
)
self.classifier = nn.Sequential(
nn.Flatten(),
nn.Linear(64, 2)
)
def forward(self, x):
x = self.features(x)
x = self.classifier(x)
return x
device = "cuda"
model = CNN().to(device)
print(model)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-3)
CNN(
(features): Sequential(
(0): Conv2d(1, 16, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): ReLU()
(2): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)
(3): Conv2d(16, 32, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): ReLU()
(5): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)
(6): Conv2d(32, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(7): ReLU()
(8): AdaptiveAvgPool2d(output_size=(1, 1))
)
(classifier): Sequential(
(0): Flatten(start_dim=1, end_dim=-1)
(1): Linear(in_features=64, out_features=2, bias=True)
)
)
5) Training loop#
def run_epoch(model, loader, criterion, optimizer=None):
if optimizer is None:
model.eval()
else:
model.train()
total_loss = 0
total_correct = 0
total_samples = 0
for x, y in loader:
x = x.to(device)
y = y.squeeze().long().to(device)
with torch.set_grad_enabled(optimizer is not None):
logits = model(x)
loss = criterion(logits, y)
if optimizer is not None:
optimizer.zero_grad()
loss.backward()
optimizer.step()
preds = torch.argmax(logits, dim=1)
total_loss += loss.item() * x.size(0)
total_correct += (preds == y).sum().item()
total_samples += x.size(0)
return total_loss / total_samples, total_correct / total_samples
num_epochs = 30
history = {
"train_loss": [],
"train_acc": [],
"val_loss": [],
"val_acc": []
}
for epoch in range(num_epochs):
train_loss, train_acc = run_epoch(model, train_loader, criterion, optimizer)
val_loss, val_acc = run_epoch(model, val_loader, criterion)
history["train_loss"].append(train_loss)
history["train_acc"].append(train_acc)
history["val_loss"].append(val_loss)
history["val_acc"].append(val_acc)
print(
f"Epoch {epoch+1}/{num_epochs} | "
f"train_loss={train_loss:.4f} | train_acc={train_acc:.4f} | "
f"val_loss={val_loss:.4f} | val_acc={val_acc:.4f}"
)
Epoch 1/30 | train_loss=0.5752 | train_acc=0.7421 | val_loss=0.5576 | val_acc=0.7424
Epoch 2/30 | train_loss=0.5262 | train_acc=0.7421 | val_loss=0.4907 | val_acc=0.7424
Epoch 3/30 | train_loss=0.4323 | train_acc=0.7789 | val_loss=0.4200 | val_acc=0.7996
Epoch 4/30 | train_loss=0.3338 | train_acc=0.8626 | val_loss=0.2975 | val_acc=0.8760
Epoch 5/30 | train_loss=0.2677 | train_acc=0.8927 | val_loss=0.2878 | val_acc=0.8721
Epoch 6/30 | train_loss=0.2516 | train_acc=0.8910 | val_loss=0.2846 | val_acc=0.8836
Epoch 7/30 | train_loss=0.2479 | train_acc=0.8985 | val_loss=0.2274 | val_acc=0.8950
Epoch 8/30 | train_loss=0.2275 | train_acc=0.9025 | val_loss=0.2776 | val_acc=0.8874
Epoch 9/30 | train_loss=0.2204 | train_acc=0.9059 | val_loss=0.2235 | val_acc=0.9046
Epoch 10/30 | train_loss=0.2076 | train_acc=0.9127 | val_loss=0.2190 | val_acc=0.9008
Epoch 11/30 | train_loss=0.2120 | train_acc=0.9121 | val_loss=0.2144 | val_acc=0.9084
Epoch 12/30 | train_loss=0.2235 | train_acc=0.9051 | val_loss=0.2042 | val_acc=0.9256
Epoch 13/30 | train_loss=0.1996 | train_acc=0.9189 | val_loss=0.2099 | val_acc=0.9160
Epoch 14/30 | train_loss=0.1967 | train_acc=0.9250 | val_loss=0.2198 | val_acc=0.9122
Epoch 15/30 | train_loss=0.1918 | train_acc=0.9216 | val_loss=0.2595 | val_acc=0.8989
Epoch 16/30 | train_loss=0.2091 | train_acc=0.9144 | val_loss=0.1950 | val_acc=0.9275
Epoch 17/30 | train_loss=0.1966 | train_acc=0.9182 | val_loss=0.1869 | val_acc=0.9275
Epoch 18/30 | train_loss=0.1785 | train_acc=0.9278 | val_loss=0.1824 | val_acc=0.9198
Epoch 19/30 | train_loss=0.1821 | train_acc=0.9261 | val_loss=0.1846 | val_acc=0.9198
Epoch 20/30 | train_loss=0.1858 | train_acc=0.9278 | val_loss=0.1785 | val_acc=0.9275
Epoch 21/30 | train_loss=0.1849 | train_acc=0.9254 | val_loss=0.1789 | val_acc=0.9256
Epoch 22/30 | train_loss=0.1767 | train_acc=0.9303 | val_loss=0.1712 | val_acc=0.9313
Epoch 23/30 | train_loss=0.1726 | train_acc=0.9308 | val_loss=0.1689 | val_acc=0.9313
Epoch 24/30 | train_loss=0.1840 | train_acc=0.9240 | val_loss=0.1716 | val_acc=0.9275
Epoch 25/30 | train_loss=0.1682 | train_acc=0.9352 | val_loss=0.1895 | val_acc=0.9256
Epoch 26/30 | train_loss=0.1634 | train_acc=0.9388 | val_loss=0.1685 | val_acc=0.9332
Epoch 27/30 | train_loss=0.1669 | train_acc=0.9356 | val_loss=0.1701 | val_acc=0.9275
Epoch 28/30 | train_loss=0.1691 | train_acc=0.9382 | val_loss=0.1650 | val_acc=0.9332
Epoch 29/30 | train_loss=0.1715 | train_acc=0.9314 | val_loss=0.1874 | val_acc=0.9256
Epoch 30/30 | train_loss=0.1796 | train_acc=0.9301 | val_loss=0.1628 | val_acc=0.9313
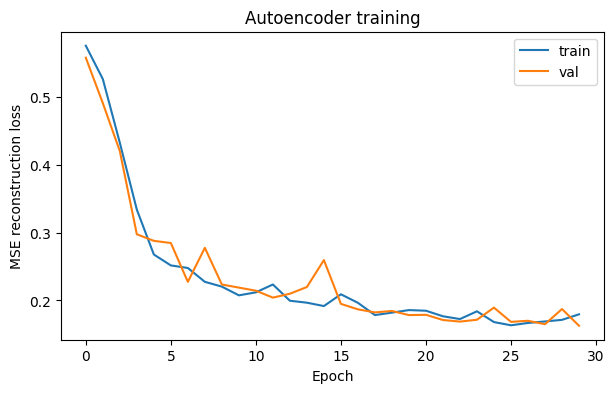
6) Plot reconstruction loss#
plt.figure(figsize=(7,4))
plt.plot(history["train_loss"], label="train")
plt.plot(history["val_loss"], label="val")
plt.xlabel("Epoch")
plt.ylabel("MSE reconstruction loss")
plt.title("Autoencoder training")
plt.legend()
plt.show()
model.eval()
all_probs = []
all_preds = []
all_targets = []
with torch.no_grad():
for x, y in test_loader:
x = x.to(device)
y = y.squeeze().long().to(device)
logits = model(x)
probs = torch.softmax(logits, dim=1)[:, 1]
preds = torch.argmax(logits, dim=1)
all_probs.extend(probs.cpu().numpy())
all_preds.extend(preds.cpu().numpy())
all_targets.extend(y.cpu().numpy())
all_probs = np.array(all_probs)
all_preds = np.array(all_preds)
all_targets = np.array(all_targets)
roc_auc = roc_auc_score(all_targets, all_probs)
fpr, tpr, thresholds = roc_curve(all_targets, all_probs)
print(f"ROC AUC: {roc_auc:.4f}")
plt.figure(figsize=(6, 6))
plt.plot(fpr, tpr, label=f"AUC = {roc_auc:.3f}")
plt.plot([0, 1], [0, 1], linestyle="--")
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
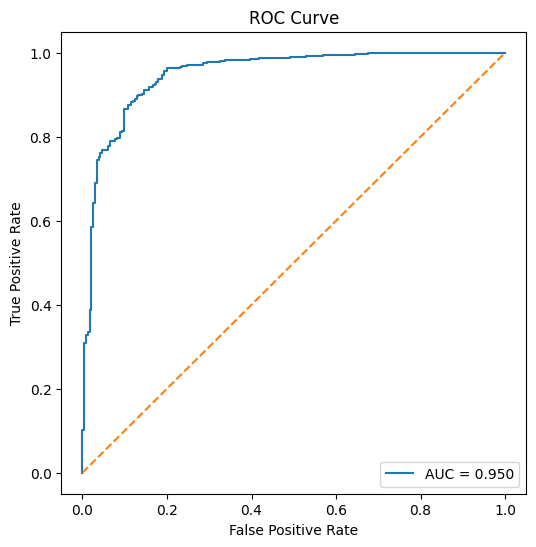
plt.title("ROC Curve")
plt.legend()
plt.show()
ROC AUC: 0.9496
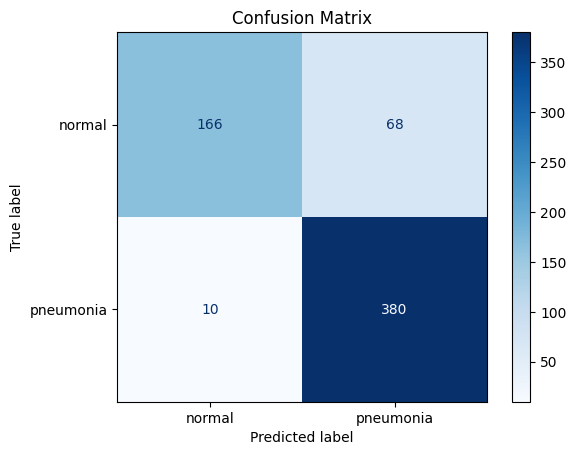
cm = confusion_matrix(all_targets, all_preds)
disp = ConfusionMatrixDisplay(
confusion_matrix=cm,
display_labels=["normal", "pneumonia"]
)
disp.plot(cmap="Blues")
plt.title("Confusion Matrix")
plt.show()
acc = accuracy_score(all_targets, all_preds)
precision = precision_score(all_targets, all_preds)
recall = recall_score(all_targets, all_preds)
f1 = f1_score(all_targets, all_preds)
print(f"Accuracy : {acc:.4f}")
print(f"Precision: {precision:.4f}")
print(f"Recall : {recall:.4f}")
print(f"F1 score : {f1:.4f}")
print(f"ROC AUC : {roc_auc:.4f}")
print("\nClassification report:\n")
print(classification_report(all_targets, all_preds, target_names=["normal", "pneumonia"]))
Accuracy : 0.8750
Precision: 0.8482
Recall : 0.9744
F1 score : 0.9069
ROC AUC : 0.9496
Classification report:
precision recall f1-score support
normal 0.94 0.71 0.81 234
pneumonia 0.85 0.97 0.91 390
accuracy 0.88 624
macro avg 0.90 0.84 0.86 624
weighted avg 0.88 0.88 0.87 624
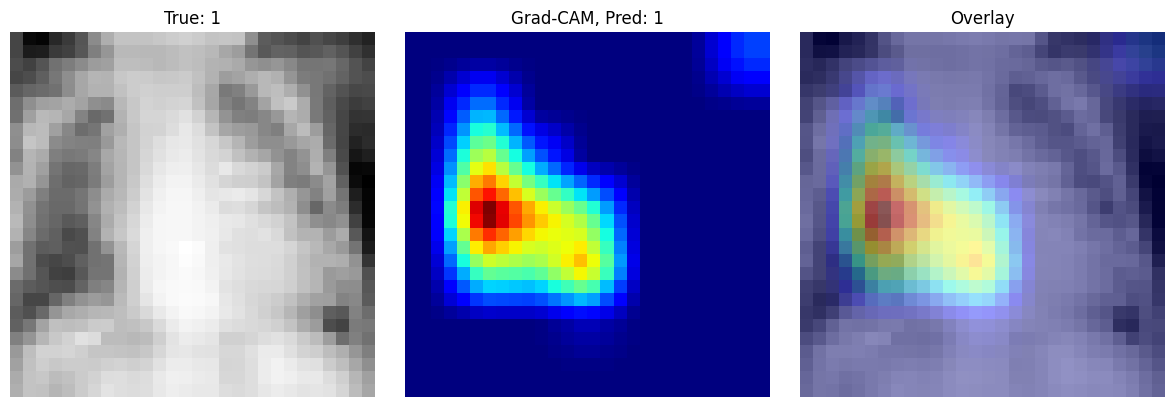
(Explainability) Grad-CAM#
Grad-CAM is an explainability method that helps us understand which image regions were most important for a model’s prediction. This can help us check whether the network is using clinically meaningful structures or whether it is relying on irrelevant patterns in the image.
The basic idea of Grad-CAM is to combine the feature maps of a convolutional layer with the gradients of the predicted class. First, we pass an image through the network and select the output score for the class of interest. Then we compute the gradient of this score with respect to the feature maps in the last convolutional layer. These gradients tell us which channels were most important for the prediction. By averaging the gradients over space, we obtain weights for each feature map. The weighted combination of these feature maps produces a coarse heatmap that highlights influential regions in the image.
class GradCAM:
def __init__(self, model, target_layer):
self.model = model
self.target_layer = target_layer
self.activations = None
self.gradients = None
self.forward_handle = target_layer.register_forward_hook(self.save_activation)
self.backward_handle = target_layer.register_full_backward_hook(self.save_gradient)
def save_activation(self, module, input, output):
self.activations = output.detach()
def save_gradient(self, module, grad_input, grad_output):
self.gradients = grad_output[0].detach()
def generate(self, x, class_idx=None):
self.model.eval()
output = self.model(x)
if class_idx is None:
class_idx = output.argmax(dim=1).item()
self.model.zero_grad()
score = output[:, class_idx]
score.backward(retain_graph=True)
weights = self.gradients.mean(dim=(2, 3), keepdim=True)
cam = (weights * self.activations).sum(dim=1, keepdim=True)
cam = F.relu(cam)
cam = F.interpolate(cam, size=x.shape[2:], mode="bilinear", align_corners=False)
cam = cam.squeeze().cpu().numpy()
cam = cam - cam.min()
cam = cam / (cam.max() + 1e-8)
return cam, class_idx
def remove_hooks(self):
self.forward_handle.remove()
self.backward_handle.remove()
target_layer = model.features[6]
gradcam = GradCAM(model, target_layer=model.features[6])
img, label = test_dataset[0]
x = img.unsqueeze(0).to(device)
cam, pred_class = gradcam.generate(x)
img_np = img.squeeze().cpu().numpy()
plt.figure(figsize=(12, 4))
plt.subplot(1, 3, 1)
plt.imshow(img_np, cmap="gray")
plt.title(f"True: {label.item()}")
plt.axis("off")
plt.subplot(1, 3, 2)
plt.imshow(cam, cmap="jet")
plt.title(f"Grad-CAM, Pred: {pred_class}")
plt.axis("off")
plt.subplot(1, 3, 3)
plt.imshow(img_np, cmap="gray")
plt.imshow(cam, cmap="jet", alpha=0.4)
plt.title("Overlay")
plt.axis("off")
plt.tight_layout()
plt.show()