Neural Network with PyTorch#
We want to build and train a so-called Multi-Layer-Perceptron (MLP). This is a modern feedforward neural network consisting of fully connected neurons with nonlinear activation functions, organized in layers.
For this, we use PyTorch, a library for deep learning using GPUs and CPUs.
The goal is to apply the MLP to a classification problem on tabular medical data (Breast Cancer Dataset).
Goal:
Load a tabular breast cancer dataset
Preprocess and split data
Build a neural network in PyTorch
Train with a standard PyTorch training loop
Evaluate performance
Experiment with architecture and hyperparameters
# Just in case we need help
# Import bia-bob as a helpful Python & Medical AI expert
from bia_bob import bob
import os
bob.initialize(
endpoint=os.getenv('ENDPOINT_URL'),
model="vllm-llama-4-scout-17b-16e-instruct",
system_prompt=os.getenv('SYSTEM_PROMPT_MEDICAL_AI')
)
%bob Who are you ? Just 1 sentence!
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import accuracy_score, classification_report, roc_auc_score, roc_curve, auc
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
from torch.utils.data import TensorDataset, DataLoader
SEED = 42
np.random.seed(SEED)
torch.manual_seed(SEED)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("Using device:", device)
Using device: cpu
1) Load breast cancer dataset#
data = load_breast_cancer(as_frame=True)
X = data.data
y = data.target
print("Features shape:", X.shape)
print("Target shape:", y.shape)
print("Classes:", data.target_names.tolist())
print("Class counts:", y.value_counts().to_dict())
Features shape: (569, 30)
Target shape: (569,)
Classes: ['malignant', 'benign']
Class counts: {1: 357, 0: 212}
display(X.head())
display(y.head())
| mean radius | mean texture | mean perimeter | mean area | mean smoothness | mean compactness | mean concavity | mean concave points | mean symmetry | mean fractal dimension | ... | worst radius | worst texture | worst perimeter | worst area | worst smoothness | worst compactness | worst concavity | worst concave points | worst symmetry | worst fractal dimension | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 17.99 | 10.38 | 122.80 | 1001.0 | 0.11840 | 0.27760 | 0.3001 | 0.14710 | 0.2419 | 0.07871 | ... | 25.38 | 17.33 | 184.60 | 2019.0 | 0.1622 | 0.6656 | 0.7119 | 0.2654 | 0.4601 | 0.11890 |
| 1 | 20.57 | 17.77 | 132.90 | 1326.0 | 0.08474 | 0.07864 | 0.0869 | 0.07017 | 0.1812 | 0.05667 | ... | 24.99 | 23.41 | 158.80 | 1956.0 | 0.1238 | 0.1866 | 0.2416 | 0.1860 | 0.2750 | 0.08902 |
| 2 | 19.69 | 21.25 | 130.00 | 1203.0 | 0.10960 | 0.15990 | 0.1974 | 0.12790 | 0.2069 | 0.05999 | ... | 23.57 | 25.53 | 152.50 | 1709.0 | 0.1444 | 0.4245 | 0.4504 | 0.2430 | 0.3613 | 0.08758 |
| 3 | 11.42 | 20.38 | 77.58 | 386.1 | 0.14250 | 0.28390 | 0.2414 | 0.10520 | 0.2597 | 0.09744 | ... | 14.91 | 26.50 | 98.87 | 567.7 | 0.2098 | 0.8663 | 0.6869 | 0.2575 | 0.6638 | 0.17300 |
| 4 | 20.29 | 14.34 | 135.10 | 1297.0 | 0.10030 | 0.13280 | 0.1980 | 0.10430 | 0.1809 | 0.05883 | ... | 22.54 | 16.67 | 152.20 | 1575.0 | 0.1374 | 0.2050 | 0.4000 | 0.1625 | 0.2364 | 0.07678 |
5 rows × 30 columns
0 0
1 0
2 0
3 0
4 0
Name: target, dtype: int64
2) Train/validation/test split#
We use:
train set: for fitting the model
validation set: for monitoring/tuning
test set: final evaluation
Terminology: Data leakage means information from the validation/test set accidentally influences training or preprocessing. This makes evaluation results look better than they really are.
X_trainval, X_test, y_trainval, y_test = train_test_split(
X, y,
test_size=0.2,
stratify=y,
random_state=SEED
)
X_train, X_val, y_train, y_val = train_test_split(
X_trainval, y_trainval,
test_size=0.2,
stratify=y_trainval,
random_state=SEED
)
print("Train:", X_train.shape, y_train.shape)
print("Val: ", X_val.shape, y_val.shape)
print("Test: ", X_test.shape, y_test.shape)
Train: (364, 30) (364,)
Val: (91, 30) (91,)
Test: (114, 30) (114,)
3) Feature scaling (important for neural networks)#
Fit scaler on training data only, then transform val/test.
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_val_scaled = scaler.transform(X_val)
X_test_scaled = scaler.transform(X_test)
y_train_np = y_train.to_numpy().astype(np.float32).reshape(-1, 1)
y_val_np = y_val.to_numpy().astype(np.float32).reshape(-1, 1)
y_test_np = y_test.to_numpy().astype(np.float32).reshape(-1, 1)
4) Convert to PyTorch tensors + DataLoaders#
X_train_tensor = torch.tensor(X_train_scaled, dtype=torch.float32)
X_val_tensor = torch.tensor(X_val_scaled, dtype=torch.float32)
X_test_tensor = torch.tensor(X_test_scaled, dtype=torch.float32)
y_train_tensor = torch.tensor(y_train_np, dtype=torch.float32)
y_val_tensor = torch.tensor(y_val_np, dtype=torch.float32)
y_test_tensor = torch.tensor(y_test_np, dtype=torch.float32)
train_ds = TensorDataset(X_train_tensor, y_train_tensor)
val_ds = TensorDataset(X_val_tensor, y_val_tensor)
test_ds = TensorDataset(X_test_tensor, y_test_tensor)
train_loader = DataLoader(train_ds, batch_size=32, shuffle=True)
val_loader = DataLoader(val_ds, batch_size=64, shuffle=False)
test_loader = DataLoader(test_ds, batch_size=64, shuffle=False)
print("Train batches:", len(train_loader))
print("Val batches:", len(val_loader))
Train batches: 12
Val batches: 2
5) Define the MLP model#
Terminology: An MLP is a neural network made of stacked fully connected layers with activation functions in between. It is commonly used for tabular data and is the easiest and most light-weight form of neural nentworks.
class NeuralNetwork(nn.Module):
def __init__(self, input_dim, hidden_dim=32, dropout=0.2):
super().__init__()
self.fc1 = nn.Linear(input_dim, hidden_dim)
self.act1 = nn.ReLU()
self.drop1 = nn.Dropout(dropout)
self.fc2 = nn.Linear(hidden_dim, hidden_dim)
self.act2 = nn.ReLU()
self.drop2 = nn.Dropout(dropout)
self.fc3 = nn.Linear(hidden_dim, 1)
def forward(self, x):
x = self.fc1(x)
x = self.act1(x)
x = self.drop1(x)
x = self.fc2(x)
x = self.act2(x)
x = self.drop2(x)
x = self.fc3(x)
return x
input_dim = X_train_tensor.shape[1]
model = NeuralNetwork(
input_dim=input_dim,
hidden_dim=32,
dropout=0.1
).to(device)
print(model)
NeuralNetwork(
(fc1): Linear(in_features=30, out_features=32, bias=True)
(act1): ReLU()
(drop1): Dropout(p=0.1, inplace=False)
(fc2): Linear(in_features=32, out_features=32, bias=True)
(act2): ReLU()
(drop2): Dropout(p=0.1, inplace=False)
(fc3): Linear(in_features=32, out_features=1, bias=True)
)
6) Define loss and optimizer#
We use:
BCEWithLogitsLossfor binary classificationAdamoptimizer for stable training
criterion = nn.BCEWithLogitsLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=1e-4)
7) Helper functions for training and evaluation#
def train_one_epoch(model, loader, criterion, optimizer, device):
model.train()
running_loss = 0.0
all_logits = []
all_targets = []
for xb, yb in loader:
xb = xb.to(device)
yb = yb.to(device)
optimizer.zero_grad()
logits = model(xb)
loss = criterion(logits, yb)
loss.backward()
optimizer.step()
running_loss += loss.item() * xb.size(0)
all_logits.append(logits.detach().cpu())
all_targets.append(yb.detach().cpu())
epoch_loss = running_loss / len(loader.dataset)
all_logits = torch.cat(all_logits).numpy()
all_targets = torch.cat(all_targets).numpy()
probs = 1 / (1 + np.exp(-all_logits)) # sigmoid to convert logits to probabilities
preds = (probs >= 0.5).astype(np.float32)
epoch_acc = accuracy_score(all_targets, preds)
return epoch_loss, epoch_acc
@torch.no_grad()
def evaluate(model, loader, criterion, device):
model.eval()
running_loss = 0.0
all_logits = []
all_targets = []
for xb, yb in loader:
xb = xb.to(device)
yb = yb.to(device)
logits = model(xb)
loss = criterion(logits, yb)
running_loss += loss.item() * xb.size(0)
all_logits.append(logits.cpu())
all_targets.append(yb.cpu())
epoch_loss = running_loss / len(loader.dataset)
all_logits = torch.cat(all_logits).numpy()
all_targets = torch.cat(all_targets).numpy()
probs = 1 / (1 + np.exp(-all_logits))
preds = (probs >= 0.5).astype(np.float32)
epoch_acc = accuracy_score(all_targets, preds)
return epoch_loss, epoch_acc, probs, preds, all_targets
8) Training loop (multiple epochs)#
num_epochs = 100
history = {
"train_loss": [],
"train_acc": [],
"val_loss": [],
"val_acc": [],
}
best_val_loss = float("inf")
best_state_dict = None
for epoch in range(1, num_epochs + 1):
train_loss, train_acc = train_one_epoch(model, train_loader, criterion, optimizer, device)
val_loss, val_acc, _, _, _ = evaluate(model, val_loader, criterion, device)
history["train_loss"].append(train_loss)
history["train_acc"].append(train_acc)
history["val_loss"].append(val_loss)
history["val_acc"].append(val_acc)
if val_loss < best_val_loss:
best_val_loss = val_loss
best_state_dict = {k: v.cpu().clone() for k, v in model.state_dict().items()}
if epoch == 1 or epoch % 10 == 0 or epoch == num_epochs:
print(
f"Epoch {epoch:03d}/{num_epochs} | "
f"train_loss={train_loss:.4f}, train_acc={train_acc:.4f} | "
f"val_loss={val_loss:.4f}, val_acc={val_acc:.4f}"
)
# Restore best validation model
if best_state_dict is not None:
model.load_state_dict(best_state_dict)
Epoch 001/100 | train_loss=0.6919, train_acc=0.4643 | val_loss=0.6873, val_acc=0.4725
Epoch 010/100 | train_loss=0.6216, train_acc=0.8571 | val_loss=0.6187, val_acc=0.9011
Epoch 020/100 | train_loss=0.5047, train_acc=0.9478 | val_loss=0.5058, val_acc=0.9341
Epoch 030/100 | train_loss=0.3729, train_acc=0.9478 | val_loss=0.3751, val_acc=0.9341
Epoch 040/100 | train_loss=0.2724, train_acc=0.9505 | val_loss=0.2791, val_acc=0.9341
Epoch 050/100 | train_loss=0.2132, train_acc=0.9560 | val_loss=0.2168, val_acc=0.9451
Epoch 060/100 | train_loss=0.1696, train_acc=0.9615 | val_loss=0.1743, val_acc=0.9560
Epoch 070/100 | train_loss=0.1384, train_acc=0.9698 | val_loss=0.1449, val_acc=0.9560
Epoch 080/100 | train_loss=0.1221, train_acc=0.9725 | val_loss=0.1241, val_acc=0.9670
Epoch 090/100 | train_loss=0.1011, train_acc=0.9753 | val_loss=0.1098, val_acc=0.9670
Epoch 100/100 | train_loss=0.0925, train_acc=0.9780 | val_loss=0.0992, val_acc=0.9780
9) Plot training curves#
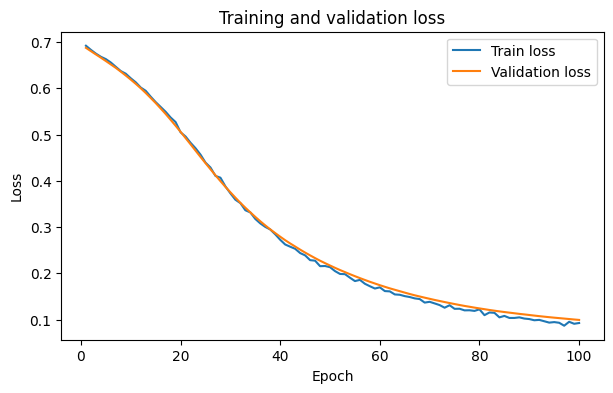
After the training we need to evaluate, how our model performed durning training. Here we have a look how the error that the model predoces behaves over the epochs of the training for the training data and the validation data.
Terminology:
Underfitting - the model is too simple to learn the pattern, so performance is poor on both training and validation/test data.
Overfitting - the model fits training data very well but does not generalize, so validation/test performance is worse.
epochs = np.arange(1, num_epochs + 1)
plt.figure(figsize=(7, 4))
plt.plot(epochs, history["train_loss"], label="Train loss")
plt.plot(epochs, history["val_loss"], label="Validation loss")
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.title("Training and validation loss")
plt.legend()
plt.show()
plt.figure(figsize=(7, 4))
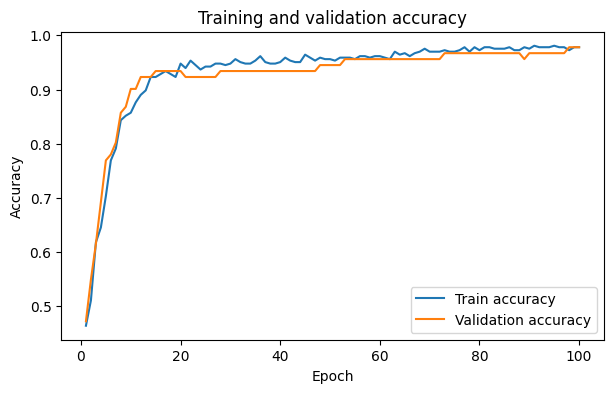
plt.plot(epochs, history["train_acc"], label="Train accuracy")
plt.plot(epochs, history["val_acc"], label="Validation accuracy")
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
plt.title("Training and validation accuracy")
plt.legend()
plt.show()
10) Final evaluation on test set#
We also need to evaluate how our model will perform on new, unseen data. Therefore, we have the test dataset. We can generate metrics derived from the prediction values of the model and the actual labels of the patients.
test_loss, test_acc, test_probs, test_preds, test_targets = evaluate(
model, test_loader, criterion, device
)
print(f"Test loss: {test_loss:.4f}")
print(f"Test accuracy: {test_acc:.4f}")
print("\nClassification report:")
print(classification_report(test_targets, test_preds, target_names=data.target_names))
Test loss: 0.1151
Test accuracy: 0.9561
Classification report:
precision recall f1-score support
malignant 0.93 0.95 0.94 42
benign 0.97 0.96 0.97 72
accuracy 0.96 114
macro avg 0.95 0.96 0.95 114
weighted avg 0.96 0.96 0.96 114
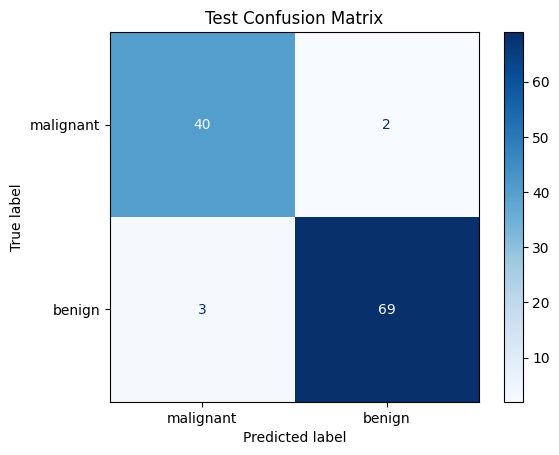
cm = confusion_matrix(test_targets, test_preds)
disp = ConfusionMatrixDisplay(confusion_matrix=cm, display_labels=data.target_names)
disp.plot(cmap="Blues", values_format="d")
plt.title("Test Confusion Matrix")
plt.show()
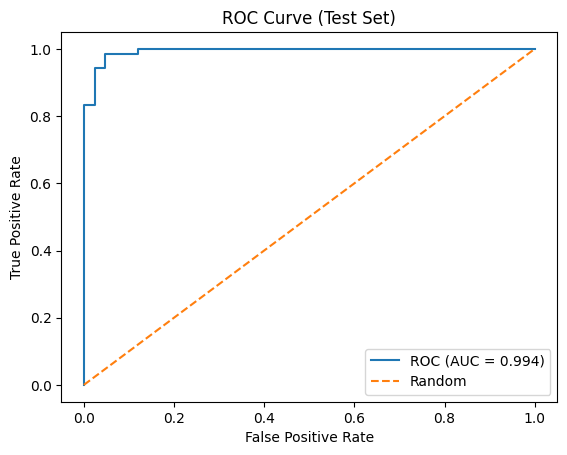
fpr, tpr, thresholds = roc_curve(test_targets.ravel(), test_probs.ravel())
roc_auc = auc(fpr, tpr)
plt.figure()
plt.plot(fpr, tpr, label=f"ROC (AUC = {roc_auc:.3f})")
plt.plot([0, 1], [0, 1], linestyle="--", label="Random")
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("ROC Curve (Test Set)")
plt.legend()
plt.show()
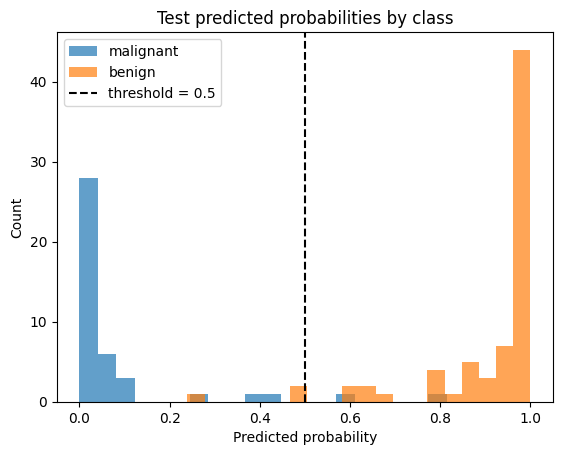
p = test_probs.ravel()
y = test_targets.ravel()
plt.figure()
plt.hist(p[y == 0], bins=20, alpha=0.7, label=f"{data.target_names[0]}")
plt.hist(p[y == 1], bins=20, alpha=0.7, label=f"{data.target_names[1]}")
plt.axvline(0.5, linestyle="--", label="threshold = 0.5", c='black')
plt.xlabel("Predicted probability")
plt.ylabel("Count")
plt.title("Test predicted probabilities by class")
plt.legend()
plt.show()
Exercise 1: What happens if you change the learning rate ? For example, try out 1e-2, 1e-6, 1e-10.
Exercise 2: Adapt the epochs also in regard to changed learning rates. When do you think your model finishes learning ?
Exercise 3: Change the network architecture. Add or remove linear layers, change dropout rates and activation functions. How does the behaviour of the model change ?
Exercise 4: (Optional) Implement you a 5-fold cross-validation training loop and see check how variable your results are across different splits of the data. (You may ask bob for help.)
