Comprehensive Exercise on Segmentation Analysis - Solution#
Spoiler:
This notebook contains a comprehensive step-by-step solution to the exercise.
Try working through the exercise on your own first before looking at this suggested solution.
One extremely helpful application of Machine Learning is in data exploration and segementation analysis. Clustering algorithms help us find fundamental structures in data that is unknown to us, e.g., whether individual data points have any similarities that we do not yet recognize.
Let’s try this out on a dataset about demographics, health metrics, and engagement patterns of patients.
We will use the Patient Segmentation Dataset from Kaggle (CC0: Public Domain).
The dataset contains 2000 patient records with comprehensive information including:
Demographics like age, gender, geographic location
Health metrics like height, weight, chronic conditions
Healthcare utilization like annual visit frequency
Insurance & engagement like insurance type, preventive care participation flag
The goal is to apply clustering on the patient data for a so-called Segmentation Analysis for identifying distinct patient personas (e.g., “High-risk seniors”, “Young healthy adults”, “Chronic disease patients”).
Your tasks:
Load the data from the file patient_segmentation_dataset.csv
Inspect the data and make yourself familiar with its content, data types, distributions, etc.
Decide for a fitting clustering algorithm based on your data insights (e.g., can KMeans be used for this specific data?)
Prepare the data for clustering and apply the clustering algorithm. Compare different clustering algorithms.
Explore the clusters (patient segments) to gain helpful insights
# Import bia-bob as a helpful Python & Medical AI expert
from bia_bob import bob
import os
bob.initialize(
endpoint=os.getenv('ENDPOINT_URL'),
model="vllm-llama-4-scout-17b-16e-instruct",
system_prompt=os.getenv('SYSTEM_PROMPT_MEDICAL_AI')
)
%bob Who are you? Just one sentence!
I’m an expert in medical data science and a skilled Python programmer and data analyst with extensive experience working with various medical datasets and applying data analysis, machine learning, and deep learning techniques.
Analysis Plan#
We approach this analysis in structured steps:
Load & inspect the data: understand shape, types, distributions, and data quality
Exploratory Data Analysis (EDA): visualize distributions and relationships
Feature selection & preprocessing: decide which features to use and how to prepare them, encode categoricals, scale numerics
Algorithm selection and cluster profiling: apply and compare different clustering algorithms, profile each cluster to identify patient personas
Visualize and compare clusters: use dimensionality reduction (PCA, t-SNE) for 2D visualization
Wrap-up: summarize findings and discuss limitations
Step 1: Load and Inspect the Data#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Load the dataset
# Notes:
# - set PatientID as dataframe index: index_col=[colum index]
# - convert columns with dates to datatime format: parse_dates=[colum index]
# - prevents pandas from interpreting the string "None" as a missing value (NaN): keep_default_na=False
df = pd.read_csv('patient_segmentation_dataset.csv', sep=',', index_col=[0], parse_dates=[13], keep_default_na=False)
df.info()
<class 'pandas.DataFrame'>
Index: 2000 entries, P10000 to P11999
Data columns (total 15 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 Age 2000 non-null int64
1 Gender 2000 non-null str
2 State 2000 non-null str
3 City 2000 non-null str
4 Height_cm 2000 non-null int64
5 Weight_kg 2000 non-null int64
6 BMI 2000 non-null float64
7 Insurance_Type 2000 non-null str
8 Primary_Condition 2000 non-null str
9 Num_Chronic_Conditions 2000 non-null int64
10 Annual_Visits 2000 non-null int64
11 Avg_Billing_Amount 2000 non-null float64
12 Last_Visit_Date 2000 non-null datetime64[us]
13 Days_Since_Last_Visit 2000 non-null int64
14 Preventive_Care_Flag 2000 non-null int64
dtypes: datetime64[us](1), float64(2), int64(7), str(5)
memory usage: 250.0+ KB
df.head(10)
| Age | Gender | State | City | Height_cm | Weight_kg | BMI | Insurance_Type | Primary_Condition | Num_Chronic_Conditions | Annual_Visits | Avg_Billing_Amount | Last_Visit_Date | Days_Since_Last_Visit | Preventive_Care_Flag | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PatientID | |||||||||||||||
| P10000 | 64 | Male | GA | Unknown | 151 | 115 | 50.4 | Private | Arthritis | 3 | 7 | 2995.0 | 2025-07-18 | 186 | 0 |
| P10001 | 59 | Male | OH | Unknown | 189 | 68 | 19.0 | Medicare | Depression | 1 | 8 | 1209.0 | 2025-12-12 | 39 | 0 |
| P10002 | 58 | Female | PA | Unknown | 156 | 91 | 37.4 | Private | Asthma | 1 | 4 | 999.0 | 2025-09-16 | 126 | 0 |
| P10003 | 43 | Female | GA | Unknown | 152 | 92 | 39.8 | Medicare | Hypertension | 1 | 6 | 5638.5 | 2025-04-09 | 286 | 1 |
| P10004 | 53 | Female | NC | Unknown | 167 | 51 | 18.3 | Medicaid | Asthma | 1 | 4 | 5796.0 | 2025-03-07 | 319 | 0 |
| P10005 | 73 | Male | OH | Unknown | 160 | 120 | 46.9 | Medicare | Anxiety | 3 | 9 | 2397.5 | 2025-05-13 | 252 | 1 |
| P10006 | 34 | Male | PA | Unknown | 161 | 51 | 19.7 | Private | None | 0 | 3 | 4668.0 | 2025-06-11 | 223 | 1 |
| P10007 | 64 | Female | MI | Unknown | 177 | 102 | 32.6 | Private | Arthritis | 3 | 1 | 9942.5 | 2025-05-13 | 252 | 0 |
| P10008 | 61 | Female | PA | Unknown | 171 | 86 | 29.4 | Private | Arthritis | 2 | 7 | 1036.0 | 2025-05-14 | 251 | 1 |
| P10009 | 34 | Male | FL | Tampa | 182 | 111 | 33.5 | Medicare | None | 0 | 1 | 4602.0 | 2026-01-09 | 11 | 0 |
# Summary statistics for numeric columns
df.describe(include=['int64', 'float64']).T
| count | mean | std | min | 25% | 50% | 75% | max | |
|---|---|---|---|---|---|---|---|---|
| Age | 2000.0 | 50.69550 | 15.444450 | 18.0 | 40.0 | 51.00 | 63.250 | 87.0 |
| Height_cm | 2000.0 | 167.90750 | 12.684494 | 145.0 | 158.0 | 168.00 | 177.000 | 195.0 |
| Weight_kg | 2000.0 | 85.14350 | 20.385428 | 50.0 | 67.0 | 86.00 | 103.000 | 120.0 |
| BMI | 2000.0 | 30.74065 | 8.839952 | 13.4 | 23.8 | 30.10 | 37.025 | 57.1 |
| Num_Chronic_Conditions | 2000.0 | 1.08000 | 0.890504 | 0.0 | 1.0 | 1.00 | 1.000 | 3.0 |
| Annual_Visits | 2000.0 | 5.46650 | 3.485965 | 1.0 | 3.0 | 4.00 | 8.000 | 12.0 |
| Avg_Billing_Amount | 2000.0 | 4000.27050 | 2463.239215 | 207.0 | 2061.0 | 3707.25 | 5650.875 | 12467.5 |
| Days_Since_Last_Visit | 2000.0 | 180.08500 | 104.688484 | 1.0 | 90.0 | 183.00 | 268.000 | 365.0 |
| Preventive_Care_Flag | 2000.0 | 0.46400 | 0.498827 | 0.0 | 0.0 | 0.00 | 1.000 | 1.0 |
# Summary statistics for categorical columns
df.describe(include='str').T
| count | unique | top | freq | |
|---|---|---|---|---|
| Gender | 2000 | 2 | Female | 1001 |
| State | 2000 | 10 | NC | 213 |
| City | 2000 | 20 | Unknown | 1012 |
| Insurance_Type | 2000 | 4 | Medicare | 906 |
| Primary_Condition | 2000 | 10 | None | 495 |
Observations from Data Inspection#
No missing values
Mixed data types: We have numerical features (Age, BMI, Avg_Billing_Amount, …), categorical features (Gender, Insurance_Type, Primary_Condition), and binary features (Preventive_Care_Flag).
Columns to potentially exclude from clustering:
Patient_ID(unique identifier),State/City(high cardinality, many ‘Unknown’),Last_Visit_Date(high cardinality, redundant withDays_Since_Last_Visit).The string
"None"inPrimary_Conditionrepresents patients without a chronic condition. This is a valid category, not a missing value.
# Collect numerical feature names
numerical_features = df.select_dtypes(include=['int64', 'float64']).columns
print('Numerical features:\n', list(numerical_features))
# Collect categorical feature names
categorical_features = [feat for feat in df.select_dtypes(include=['str']).columns if feat not in ['State', 'City']]
print('Categorical features:\n', list(categorical_features))
Numerical features:
['Age', 'Height_cm', 'Weight_kg', 'BMI', 'Num_Chronic_Conditions', 'Annual_Visits', 'Avg_Billing_Amount', 'Days_Since_Last_Visit', 'Preventive_Care_Flag']
Categorical features:
['Gender', 'Insurance_Type', 'Primary_Condition']
Step 2: Exploratory Data Analysis (EDA)#
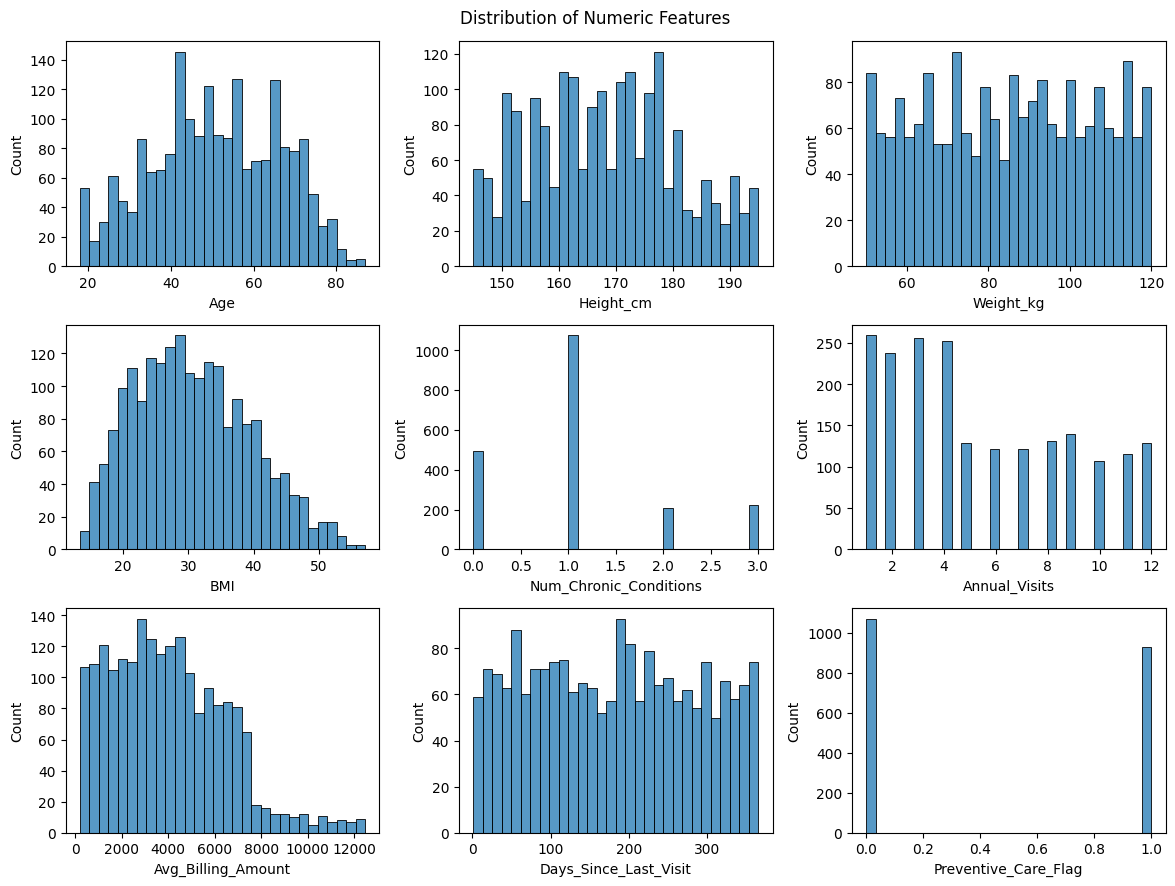
# Distribution of numerical features using histogram plots
fig, axes = plt.subplots(3, 3, figsize=(12, 9))
for ax, col in zip(axes.flatten(), numerical_features):
sns.histplot(data=df, x=col, bins=30, ax=ax)
plt.suptitle('Distribution of Numeric Features', fontsize=12)
plt.tight_layout()
plt.show()
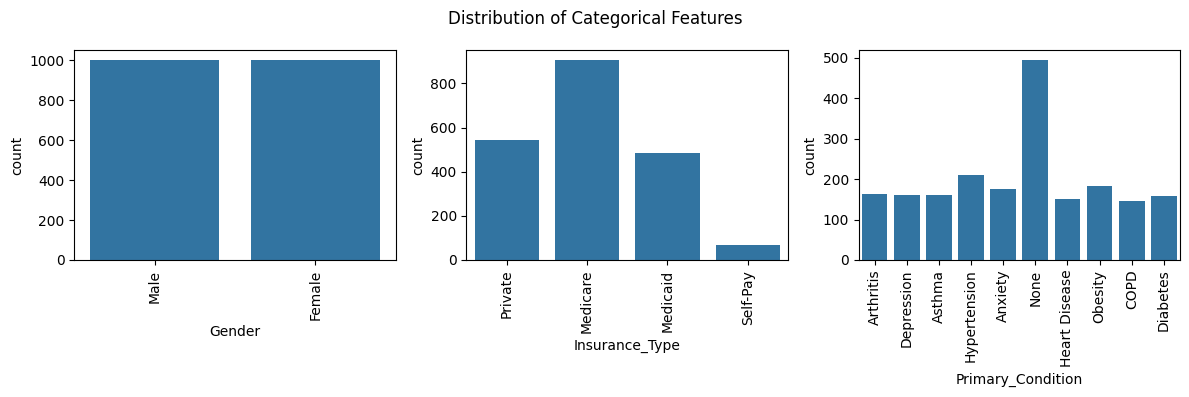
# Distribution of categorical features using count plots
fig, axes = plt.subplots(1, 3, figsize=(12, 4))
for ax, col in zip(axes.flatten(), categorical_features):
sns.countplot(data=df, x=col, ax=ax)
ax.tick_params(axis='x', rotation=90)
plt.suptitle('Distribution of Categorical Features', fontsize=12)
plt.tight_layout()
plt.show()
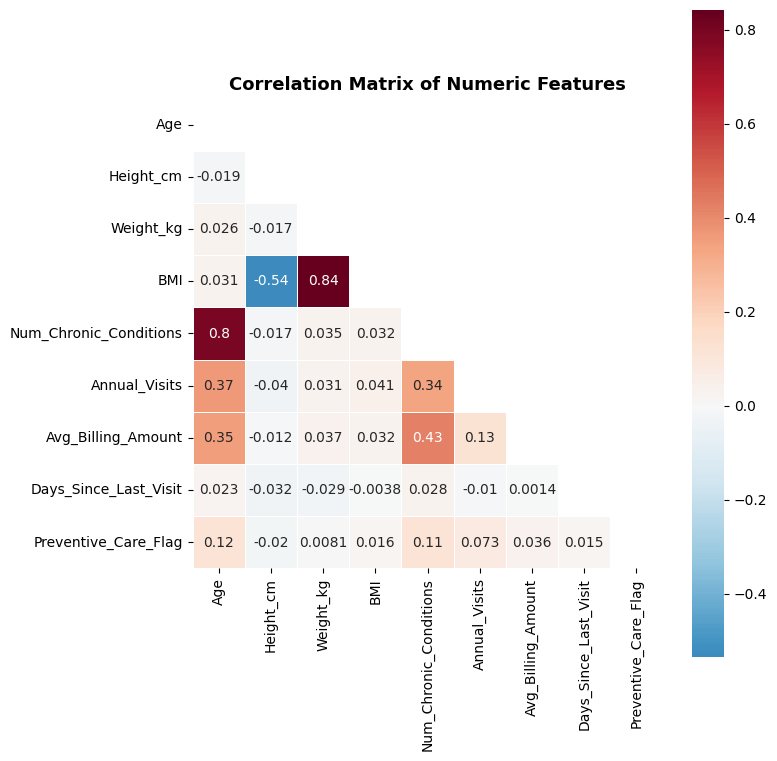
# Correlation matrix for numerical features
fig, ax = plt.subplots(figsize=(8, 8))
corr = df[numerical_features].corr()
mask = np.triu(np.ones_like(corr, dtype=bool))
sns.heatmap(corr, mask=mask, annot=True, cmap='RdBu_r', center=0, square=True, linewidths=0.5, ax=ax)
ax.set_title('Correlation Matrix of Numeric Features', fontsize=13, fontweight='bold')
plt.tight_layout()
plt.show()
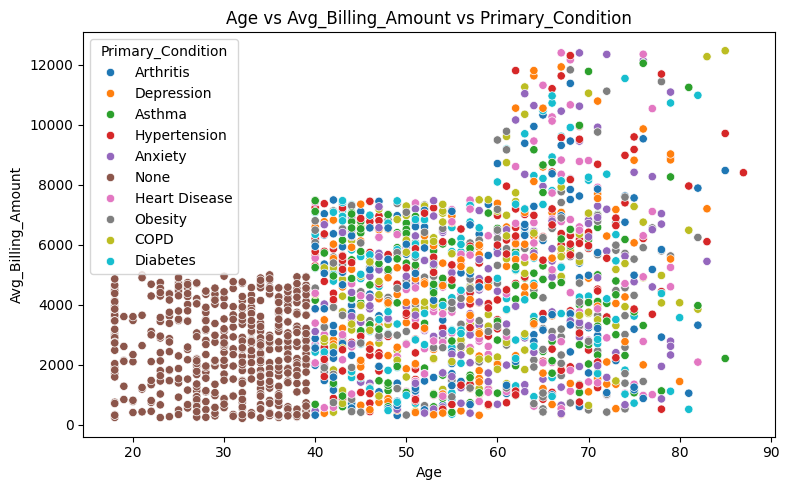
# Plot relationships: Age vs. Avg Billing Amount, colored by Primary Condition
fig, axes = plt.subplots(figsize=(8, 5))
sns.scatterplot(data=df, x='Age', y='Avg_Billing_Amount', hue='Primary_Condition', ax=axes)
axes.set_title('Age vs Avg_Billing_Amount vs Primary_Condition')
plt.tight_layout()
plt.show()
# Plot relationships: Age vs. Avg Billing Amount, colored by Insurance Type
fig, axes = plt.subplots(figsize=(8, 5))
sns.scatterplot(data=df, x='Age', y='Avg_Billing_Amount', hue='Insurance_Type', ax=axes)
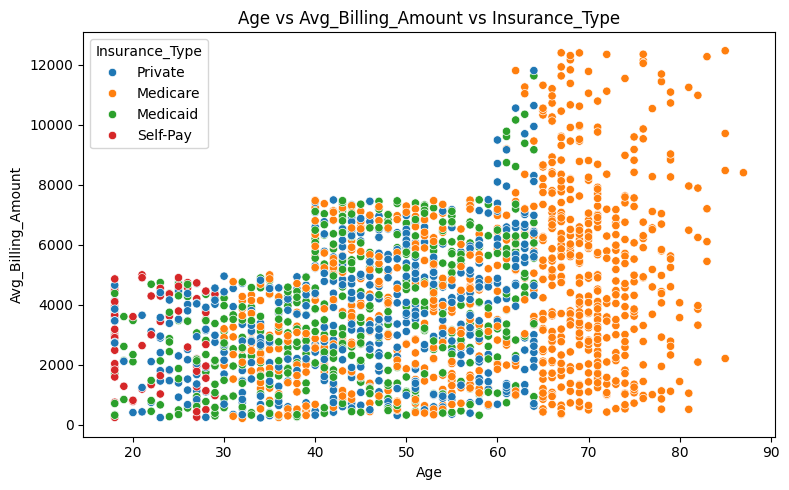
axes.set_title('Age vs Avg_Billing_Amount vs Insurance_Type')
plt.tight_layout()
plt.show()
Key EDA Insights#
Age and Gender are roughly uniformly distributed
Age, Num_Chronic_Conditions, Anual_Visits, and Avg_Billing_Amount are positively correlated – older patients visit more, have primary conditions, and cost more.
BMI is derived from Height and Weight, so including all three would introduce redundancy. We should consider dropping Height and Weight, or BMI.
Patients under 40 do not report any primary condition
Younger patients under the age of 30 are more likely to pay out of pocket, while patients over the age of 65 are insured exclusively through Medicare.
Step 3: Feature Selection and Preprocessing#
Feature decisions#
Include:
Age– key demographicBMI– derived health metric (we dropHeight_cmandWeight_kgto avoid redundancy)Num_Chronic_Conditions– health complexity indicatorAnnual_Visits– healthcare utilizationAvg_Billing_Amount– healthcare expenditureDays_Since_Last_Visit– recency of engagementPreventive_Care_Flag– engagement indicator (binary categorical with 0 / 1)Gender– key demographic (categorical, should be one-hot encoded)Insurance_Type– access indicator (categorical, should be one-hot encoded)Primary_Condition– health status (categorical, should be one-hot encoded)
Exclude:
Patient_ID– identifier, not a featureState,City– high cardinality geographic data with many unknownsLast_Visit_Date– redundant withDays_Since_Last_VisitHeight_cm,Weight_kg– redundant withBMI
# Select features for clustering
# Remove 'Height_cm' and 'Weight_kg' from numeric features
numerical_features_clustering = [feat for feat in numerical_features if feat not in ['Height_cm', 'Weight_kg']]
categorical_features_clustering = categorical_features
# Create a subset with selected features
df_features = df[numerical_features_clustering + categorical_features_clustering].copy()
print(f"Selected features for clustering: {len(df_features.columns)} columns")
df_features.head()
Selected features for clustering: 10 columns
| Age | BMI | Num_Chronic_Conditions | Annual_Visits | Avg_Billing_Amount | Days_Since_Last_Visit | Preventive_Care_Flag | Gender | Insurance_Type | Primary_Condition | |
|---|---|---|---|---|---|---|---|---|---|---|
| PatientID | ||||||||||
| P10000 | 64 | 50.4 | 3 | 7 | 2995.0 | 186 | 0 | Male | Private | Arthritis |
| P10001 | 59 | 19.0 | 1 | 8 | 1209.0 | 39 | 0 | Male | Medicare | Depression |
| P10002 | 58 | 37.4 | 1 | 4 | 999.0 | 126 | 0 | Female | Private | Asthma |
| P10003 | 43 | 39.8 | 1 | 6 | 5638.5 | 286 | 1 | Female | Medicare | Hypertension |
| P10004 | 53 | 18.3 | 1 | 4 | 5796.0 | 319 | 0 | Female | Medicaid | Asthma |
# One-hot encoding for categorical features
df_encoded = pd.get_dummies(df_features, columns=categorical_features_clustering, drop_first=False)
print(f"Shape after encoding: {df_encoded.shape}")
print(f"\nEncoded feature names:")
for i, col in enumerate(df_encoded.columns):
print(f" {i+1:2d}. {col}")
df_encoded.head()
Shape after encoding: (2000, 23)
Encoded feature names:
1. Age
2. BMI
3. Num_Chronic_Conditions
4. Annual_Visits
5. Avg_Billing_Amount
6. Days_Since_Last_Visit
7. Preventive_Care_Flag
8. Gender_Female
9. Gender_Male
10. Insurance_Type_Medicaid
11. Insurance_Type_Medicare
12. Insurance_Type_Private
13. Insurance_Type_Self-Pay
14. Primary_Condition_Anxiety
15. Primary_Condition_Arthritis
16. Primary_Condition_Asthma
17. Primary_Condition_COPD
18. Primary_Condition_Depression
19. Primary_Condition_Diabetes
20. Primary_Condition_Heart Disease
21. Primary_Condition_Hypertension
22. Primary_Condition_None
23. Primary_Condition_Obesity
| Age | BMI | Num_Chronic_Conditions | Annual_Visits | Avg_Billing_Amount | Days_Since_Last_Visit | Preventive_Care_Flag | Gender_Female | Gender_Male | Insurance_Type_Medicaid | ... | Primary_Condition_Anxiety | Primary_Condition_Arthritis | Primary_Condition_Asthma | Primary_Condition_COPD | Primary_Condition_Depression | Primary_Condition_Diabetes | Primary_Condition_Heart Disease | Primary_Condition_Hypertension | Primary_Condition_None | Primary_Condition_Obesity | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PatientID | |||||||||||||||||||||
| P10000 | 64 | 50.4 | 3 | 7 | 2995.0 | 186 | 0 | False | True | False | ... | False | True | False | False | False | False | False | False | False | False |
| P10001 | 59 | 19.0 | 1 | 8 | 1209.0 | 39 | 0 | False | True | False | ... | False | False | False | False | True | False | False | False | False | False |
| P10002 | 58 | 37.4 | 1 | 4 | 999.0 | 126 | 0 | True | False | False | ... | False | False | True | False | False | False | False | False | False | False |
| P10003 | 43 | 39.8 | 1 | 6 | 5638.5 | 286 | 1 | True | False | False | ... | False | False | False | False | False | False | False | True | False | False |
| P10004 | 53 | 18.3 | 1 | 4 | 5796.0 | 319 | 0 | True | False | True | ... | False | False | True | False | False | False | False | False | False | False |
5 rows × 23 columns
Why Scaling?#
Some clustering algorithms use distances, e.g. KMeans uses Euclidean distance. Without scaling, features with large ranges (e.g., Avg_Billing_Amount) would dominate the distance calculation over features with small ranges (e.g., Preventive_Care_Flag being 0 or 1). StandardScaler transforms each feature to have mean=0 and std=1, giving all features equal influence.
# Standardize all features
from sklearn.preprocessing import StandardScaler
print("Statistics of first 5 features before scaling:\n",
df_encoded.iloc[:, :5].describe().T[['mean', 'std']].round(4))
scaler = StandardScaler()
X_scaled = scaler.fit_transform(df_encoded)
print("\nStatistics of first 5 features after scaling:\n",
pd.DataFrame(X_scaled, columns=df_encoded.columns).iloc[:, :5].describe().T[['mean', 'std']].round(4))
Statistics of first 5 features before scaling:
mean std
Age 50.6955 15.4445
BMI 30.7406 8.8400
Num_Chronic_Conditions 1.0800 0.8905
Annual_Visits 5.4665 3.4860
Avg_Billing_Amount 4000.2705 2463.2392
Statistics of first 5 features after scaling:
mean std
Age -0.0 1.0003
BMI -0.0 1.0003
Num_Chronic_Conditions -0.0 1.0003
Annual_Visits -0.0 1.0003
Avg_Billing_Amount -0.0 1.0003
Step 4: Algorithm Selection#
Overview of Clustering Algorithms#
We apply and compare three clustering algorithms from scikit-learn on the same preprocessed and scaled patient data:
Algorithm |
Type |
Requires K? |
Handles Noise? |
Shape Assumption |
Key Hyperparameter(s) |
|---|---|---|---|---|---|
KMeans |
Centroid-based |
Yes |
No - all points assigned |
Spherical, equal-size |
|
DBSCAN |
Density-based |
Auto-discovered |
Yes - noise labeled -1 |
Arbitrary |
|
MeanShift |
Kernel density-based |
Auto-discovered |
No - all points assigned |
Arbitrary |
|
How each algorithm works:
KMeans: Assigns each point to the nearest centroid and iterates until convergence. Fast and interpretable, but requires specifying K and assumes convex, equally-sized clusters, and euclidean feature space.
DBSCAN: Identifies clusters as dense regions separated by sparser areas. Points in low-density zones are labeled noise. Powerful for irregularly shaped clusters, but sensitive to
epsandmin_samples.MeanShift: Shifts a kernel window towards local density maxima. Discovers K automatically based on data density. The
bandwidthparameter controls kernel size.
The following steps on clustering are:
Step 4.1: KMeans
Elbow method + silhouette analysis to select K
Fit KMeans with K
Numeric profile table + detailed per-cluster summaries + heatmap
Step 4.2: DBSCAN
k-distance plot for eps selection
Fit DBSCAN
Numeric profile table + detailed per-cluster summaries + heatmap
Step 4.3: MeanShift
Bandwidth estimation with estimate_bandwidth
Fit MeanShift
Numeric profile table + detailed per-cluster summaries + heatmap
Step 5 - Side-by-side comparison
PCA projection (1×3 subplots: KMeans | DBSCAN | MeanShift), with KMeans cluster centers marked
t-SNE projection (same layout)
Quantitative metrics table (Silhouette, Davies-Bouldin, Inertia)
After fitting all three algorithms, we compare their results using PCA and t-SNE projections in Step 5.
Here, we define some more methods for further cluster analysis:
profile_cols = ['Age', 'BMI', 'Num_Chronic_Conditions', 'Annual_Visits', 'Avg_Billing_Amount', 'Days_Since_Last_Visit']
# Method for printing descriptive statistics for a specified cluster algorithm
def print_cluster_profile(clustering='Cluster_KMeans'):
cluster_profile = df.groupby(clustering)[profile_cols].agg(['mean', 'std']).round(2)
display(cluster_profile)
# Method for printing aggregated statistics for a specified cluster algorithm and cluster number
def print_cluster_statistics(clustering='Cluster_KMeans', cluster_nb=0):
grp = df[df[clustering] == cluster_nb]
n = len(grp)
print(f"\n{'='*70}")
print(f" CLUSTER {cluster_nb} (n={n}, {n/len(df)*100:.1f}% of patients)")
print(f"{'='*70}")
print(f" Age: {grp['Age'].mean():.1f} ± {grp['Age'].std():.1f}")
print(f" BMI: {grp['BMI'].mean():.1f} ± {grp['BMI'].std():.1f}")
print(f" Chronic conditions: {grp['Num_Chronic_Conditions'].mean():.2f} (avg)")
top_cond = grp['Primary_Condition'].value_counts().head(3)
print(f" Top conditions: {', '.join([f'{cd} ({ct})' for cd, ct in top_cond.items()])}")
print(f" Annual visits: {grp['Annual_Visits'].mean():.1f}")
print(f" Avg billing: ${grp['Avg_Billing_Amount'].mean():,.0f}")
print(f" Preventive care: {grp['Preventive_Care_Flag'].mean()*100:.1f}%")
gender_pct = grp['Gender'].value_counts(normalize=True) * 100
print(f" Gender: {', '.join([f'{g}: {p:.0f}%' for g, p in gender_pct.items()])}")
ins_pct = grp['Insurance_Type'].value_counts(normalize=True) * 100
print(f" Insurance: {', '.join([f'{t}: {p:.0f}%' for t, p in ins_pct.items()])}")
# Method for visualizing cluster means
def plot_cluster_means_heatmap(clustering='Cluster_KMeans'):
cluster_means = df.groupby(clustering)[profile_cols].mean()
norm = (cluster_means - cluster_means.min()) / (cluster_means.max() - cluster_means.min())
fig, ax = plt.subplots(figsize=(10, 4))
sns.heatmap(norm.T, annot=cluster_means.T.round(1).values, fmt='', annot_kws={"size": 8},
cmap='YlOrRd', linewidths=1, ax=ax, cbar_kws={'label': 'Normalized value'})
ax.set_xlabel(clustering, fontsize=10)
ax.set_title('Cluster Profiles Comparison\n(numbers = actual means, color = normalized)', fontsize=12,)
plt.tight_layout()
plt.show()
Step 4.1: KMeans#
KMeans partitions data into K clusters by minimizing within-cluster variance (inertia). Each point is assigned to its nearest centroid; centroids are updated iteratively until convergence.
Assumptions:
All features are numeric and on a comparable scale (met via StandardScaler)
Clusters are roughly spherical and similarly sized
K must be specified in advance, we determine it with the methods below
Determine the Optimal Number of Clusters#
We use two complementary methods:
Elbow Method - plot inertia (within-cluster sum of squares) vs. K; look for the “elbow” where gains diminish
Silhouette Score - measures how well each point fits its own cluster vs. its nearest neighbor cluster (range −1 to 1, higher is better)
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
K_range = range(2, 15)
inertias = []
silhouette_scores_kmeans = []
for k in K_range:
km = KMeans(n_clusters=k, random_state=42, n_init=10)
labels = km.fit_predict(X_scaled)
inertias.append(km.inertia_)
sil = silhouette_score(X_scaled, labels)
silhouette_scores_kmeans.append(sil)
print(f"K={k:2d} | Inertia: {km.inertia_:10.0f} | Silhouette: {sil:.4f}")
K= 2 | Inertia: 40206 | Silhouette: 0.1179
K= 3 | Inertia: 37111 | Silhouette: 0.1254
K= 4 | Inertia: 35237 | Silhouette: 0.1463
K= 5 | Inertia: 33327 | Silhouette: 0.1509
K= 6 | Inertia: 31437 | Silhouette: 0.2034
K= 7 | Inertia: 29238 | Silhouette: 0.2301
K= 8 | Inertia: 27051 | Silhouette: 0.2536
K= 9 | Inertia: 24850 | Silhouette: 0.2829
K=10 | Inertia: 22943 | Silhouette: 0.2929
K=11 | Inertia: 21646 | Silhouette: 0.2675
K=12 | Inertia: 19959 | Silhouette: 0.2702
K=13 | Inertia: 19342 | Silhouette: 0.2678
K=14 | Inertia: 18996 | Silhouette: 0.2578
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
# Elbow plot
ax1.plot(list(K_range), inertias, 'bs-', linewidth=1, markersize=4)
ax1.set_xlabel('Number of Clusters (K)', fontsize=12)
ax1.set_ylabel('Inertia (Within-Cluster Sum of Squares)', fontsize=12)
ax1.set_title('Elbow Method', fontsize=13, fontweight='bold')
ax1.set_xticks(list(K_range))
ax1.grid(True, alpha=0.3)
# Silhouette plot
ax2.plot(list(K_range), silhouette_scores_kmeans, 'rs-', linewidth=1, markersize=4)
ax2.set_xlabel('Number of Clusters (K)', fontsize=12)
ax2.set_ylabel('Silhouette Score', fontsize=12)
ax2.set_title('Silhouette Analysis', fontsize=13, fontweight='bold')
ax2.set_xticks(list(K_range))
ax2.grid(True, alpha=0.3)
best_k = list(K_range)[np.argmax(silhouette_scores_kmeans)]
ax2.axvline(x=best_k, color='green', linestyle='--', alpha=0.7, label=f'Best K={best_k}')
ax2.legend(fontsize=11)
plt.suptitle('KMeans: Selecting the Number of Clusters', fontsize=14, fontweight='bold')
plt.tight_layout()
plt.show()
print(f"\nBest K by Silhouette Score: {best_k} (score: {max(silhouette_scores_kmeans):.4f})")
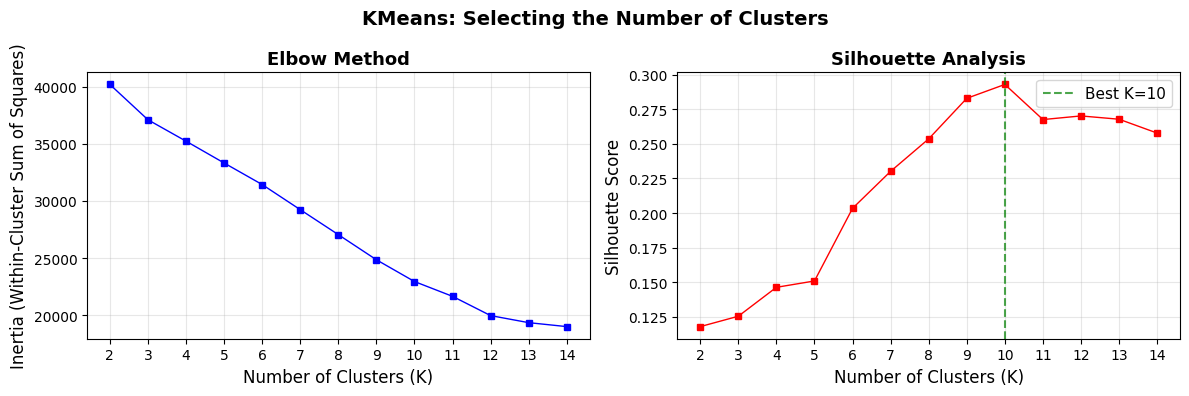
Best K by Silhouette Score: 10 (score: 0.2929)
Choosing K#
The silhouette scores are relatively low overall. This is common for real-world, high-dimensional, mixed-type data. The elbow plot unfortunatelly shows no noticeable flattening.
In practice, domain knowledge should also inform the final choice of K.
We choose K=6 as a practical balance:
Shows the first big bump in silhouette score
Provides enough granularity to identify distinct patient personas
Keeps segments manageable for clinical interpretation
Consistent with healthcare domain expectations (3–6 personas are typical)
Fit KMeans with K#
K = 6
kmeans_final = KMeans(n_clusters=K, random_state=42, n_init=10)
kmeans_labels = kmeans_final.fit_predict(X_scaled)
df['Cluster_KMeans'] = kmeans_labels
print(f"KMeans (K={K}) – Cluster sizes:")
print(df['Cluster_KMeans'].value_counts().sort_index())
print(f"\nSilhouette Score: {silhouette_score(X_scaled, kmeans_labels):.4f}")
KMeans (K=6) – Cluster sizes:
Cluster_KMeans
0 664
1 158
2 495
3 370
4 150
5 163
Name: count, dtype: int64
Silhouette Score: 0.2034
Profile KMeans Clusters#
We inspect the cluster profiles to understand the characteristics of each patient segment - looking at demographics, health metrics, and healthcare utilization.
# Print cluster profile
print_cluster_profile('Cluster_KMeans')
| Age | BMI | Num_Chronic_Conditions | Annual_Visits | Avg_Billing_Amount | Days_Since_Last_Visit | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| mean | std | mean | std | mean | std | mean | std | mean | std | mean | std | |
| Cluster_KMeans | ||||||||||||
| 0 | 57.64 | 11.18 | 30.97 | 8.97 | 1.45 | 0.75 | 6.57 | 3.40 | 4420.68 | 2466.95 | 177.27 | 106.92 |
| 1 | 56.37 | 10.79 | 31.85 | 7.86 | 1.43 | 0.73 | 6.58 | 3.49 | 4679.84 | 2665.75 | 198.32 | 102.21 |
| 2 | 30.40 | 6.39 | 29.89 | 8.83 | 0.00 | 0.00 | 2.51 | 1.09 | 2598.31 | 1393.22 | 173.49 | 103.66 |
| 3 | 57.32 | 10.97 | 31.47 | 9.08 | 1.41 | 0.72 | 6.55 | 3.47 | 4526.89 | 2618.55 | 181.07 | 104.42 |
| 4 | 58.35 | 10.77 | 30.14 | 8.47 | 1.47 | 0.77 | 5.89 | 3.65 | 4428.27 | 2720.12 | 177.96 | 99.78 |
| 5 | 56.45 | 11.74 | 30.21 | 8.85 | 1.42 | 0.73 | 6.00 | 3.38 | 4297.17 | 2581.05 | 193.61 | 104.36 |
# Print statistics for cluster n
print_cluster_statistics('Cluster_KMeans', 0)
======================================================================
CLUSTER 0 (n=664, 33.2% of patients)
======================================================================
Age: 57.6 ± 11.2
BMI: 31.0 ± 9.0
Chronic conditions: 1.45 (avg)
Top conditions: Obesity (183), Anxiety (175), Asthma (160)
Annual visits: 6.6
Avg billing: $4,421
Preventive care: 51.8%
Gender: Female: 51%, Male: 49%
Insurance: Medicare: 56%, Private: 25%, Medicaid: 18%
plot_cluster_means_heatmap('Cluster_KMeans')
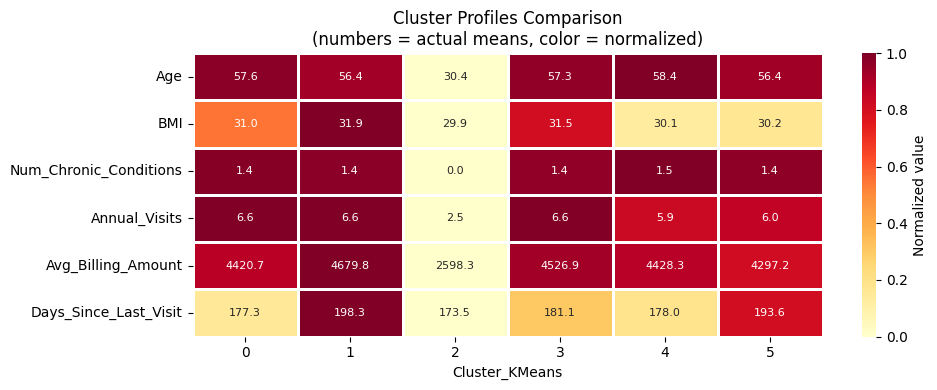
Step 4.2: DBSCAN#
DBSCAN (Density-Based Spatial Clustering of Applications with Noise) defines clusters as dense regions separated by sparser areas.
Key properties:
No need to specify K, the number of clusters is discovered automatically
Points in low-density regions are labeled as noise (label = −1) and excluded from clusters
Can find clusters of arbitrary shape
Sensitive to two hyperparameters:
eps: neighborhood radius - how far apart two points can be to be considered neighborsmin_samples: minimum number of neighbors for a point to be a core point
Select eps with the k-Distance Plot#
For each point, we compute the distance to its k-th nearest neighbor (where k = min_samples).
Sorting and plotting these distances reveals an “elbow”, the distance at the elbow is a good candidate for eps.
from sklearn.neighbors import NearestNeighbors
min_samples_dbscan = 10
nbrs = NearestNeighbors(n_neighbors=min_samples_dbscan).fit(X_scaled)
distances, _ = nbrs.kneighbors(X_scaled)
k_distances = np.sort(distances[:, -1])
plt.figure(figsize=(8, 4))
sns.lineplot(x=np.arange(len(k_distances)), y=k_distances)
plt.xlabel('Points sorted by distance', fontsize=10)
plt.ylabel(f'{min_samples_dbscan}-NN distance', fontsize=10)
plt.title('k-Distance Plot for DBSCAN eps Selection', fontsize=13, fontweight='bold')
plt.tight_layout()
plt.show()
print(f"min_samples = {min_samples_dbscan}")
print(f"Suggested eps range (80th–90th percentile of k-distances): "
f"{np.percentile(k_distances, 80):.2f} – {np.percentile(k_distances, 90):.2f}")
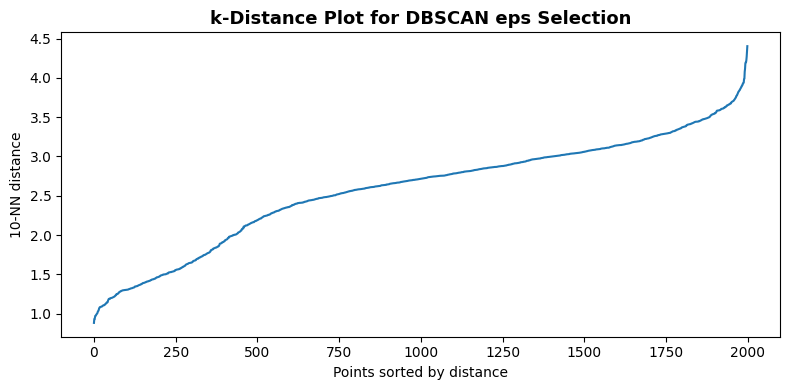
min_samples = 10
Suggested eps range (80th–90th percentile of k-distances): 3.14 – 3.37
Fit DBSCAN#
We set eps based on the elbow in the k-distance plot above. A value near the 80th–90th percentile of k-distances is a common starting point. Adjust eps up to merge clusters or down to split them.
from sklearn.cluster import DBSCAN
eps_dbscan = 3.5
dbscan = DBSCAN(eps=eps_dbscan, min_samples=min_samples_dbscan)
dbscan_labels = dbscan.fit_predict(X_scaled)
df['Cluster_DBSCAN'] = dbscan_labels
n_clusters_dbscan = len(set(dbscan_labels)) - (1 if -1 in dbscan_labels else 0)
n_noise = int((dbscan_labels == -1).sum())
print(f"DBSCAN (eps={eps_dbscan}, min_samples={min_samples_dbscan})")
print(f"Clusters found: {n_clusters_dbscan}")
print(f"Noise points: {n_noise} ({n_noise / len(dbscan_labels) * 100:.1f}%)")
print(f"\nCluster sizes:")
for label in sorted(set(dbscan_labels)):
count = (dbscan_labels == label).sum()
name = f"Cluster {label}" if label != -1 else "Noise (-1)"
print(f" {name}: {count} ({count / len(dbscan_labels) * 100:.1f}%)")
mask_non_noise = dbscan_labels != -1
if n_clusters_dbscan > 1 and mask_non_noise.sum() > 1:
sil_dbscan = silhouette_score(X_scaled[mask_non_noise], dbscan_labels[mask_non_noise])
print(f"\nSilhouette Score (non-noise points): {sil_dbscan:.4f}")
DBSCAN (eps=3.5, min_samples=10)
Clusters found: 11
Noise points: 4 (0.2%)
Cluster sizes:
Noise (-1): 4 (0.2%)
Cluster 0: 158 (7.9%)
Cluster 1: 160 (8.0%)
Cluster 2: 210 (10.5%)
Cluster 3: 174 (8.7%)
Cluster 4: 429 (21.4%)
Cluster 5: 163 (8.2%)
Cluster 6: 150 (7.5%)
Cluster 7: 183 (9.2%)
Cluster 8: 145 (7.2%)
Cluster 9: 66 (3.3%)
Cluster 10: 158 (7.9%)
Silhouette Score (non-noise points): 0.3233
Profile DBSCAN Clusters#
We inspect the cluster profiles to understand the characteristics of each patient segment - looking at demographics, health metrics, and healthcare utilization.
# Print cluster profile
print_cluster_profile('Cluster_DBSCAN')
| Age | BMI | Num_Chronic_Conditions | Annual_Visits | Avg_Billing_Amount | Days_Since_Last_Visit | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| mean | std | mean | std | mean | std | mean | std | mean | std | mean | std | |
| Cluster_DBSCAN | ||||||||||||
| -1 | 63.25 | 0.96 | 22.65 | 7.31 | 3.00 | 0.00 | 4.25 | 5.19 | 10985.00 | 852.37 | 171.75 | 150.51 |
| 0 | 57.54 | 10.63 | 31.72 | 8.77 | 1.39 | 0.67 | 6.77 | 3.29 | 4219.30 | 2559.14 | 177.58 | 102.48 |
| 1 | 57.24 | 11.19 | 32.00 | 9.55 | 1.41 | 0.72 | 6.91 | 3.34 | 4341.62 | 2399.63 | 183.83 | 103.24 |
| 2 | 57.09 | 11.28 | 31.35 | 9.31 | 1.41 | 0.74 | 6.43 | 3.59 | 4689.83 | 2567.73 | 182.60 | 105.89 |
| 3 | 58.78 | 10.95 | 31.70 | 8.79 | 1.47 | 0.77 | 6.41 | 3.46 | 4537.60 | 2637.18 | 176.84 | 112.69 |
| 4 | 31.54 | 5.92 | 29.82 | 8.81 | 0.00 | 0.00 | 2.46 | 1.10 | 2578.62 | 1381.46 | 174.38 | 103.87 |
| 5 | 56.45 | 11.74 | 30.21 | 8.85 | 1.42 | 0.73 | 6.00 | 3.38 | 4297.17 | 2581.05 | 193.61 | 104.36 |
| 6 | 58.35 | 10.77 | 30.14 | 8.47 | 1.47 | 0.77 | 5.89 | 3.65 | 4428.27 | 2720.12 | 177.96 | 99.78 |
| 7 | 57.62 | 11.53 | 29.81 | 8.48 | 1.48 | 0.76 | 6.65 | 3.37 | 4193.49 | 2385.80 | 175.57 | 103.49 |
| 8 | 56.66 | 11.07 | 30.58 | 8.98 | 1.40 | 0.73 | 6.30 | 3.38 | 4573.91 | 2351.09 | 174.48 | 108.41 |
| 9 | 23.03 | 4.06 | 30.33 | 9.02 | 0.00 | 0.00 | 2.85 | 0.98 | 2726.32 | 1471.93 | 167.76 | 102.86 |
| 10 | 56.37 | 10.79 | 31.85 | 7.86 | 1.43 | 0.73 | 6.58 | 3.49 | 4679.84 | 2665.75 | 198.32 | 102.21 |
# Print statistics for cluster n
print_cluster_statistics('Cluster_DBSCAN', 0)
======================================================================
CLUSTER 0 (n=158, 7.9% of patients)
======================================================================
Age: 57.5 ± 10.6
BMI: 31.7 ± 8.8
Chronic conditions: 1.39 (avg)
Top conditions: Depression (158)
Annual visits: 6.8
Avg billing: $4,219
Preventive care: 48.1%
Gender: Female: 54%, Male: 46%
Insurance: Medicare: 46%, Medicaid: 27%, Private: 27%
plot_cluster_means_heatmap('Cluster_DBSCAN')
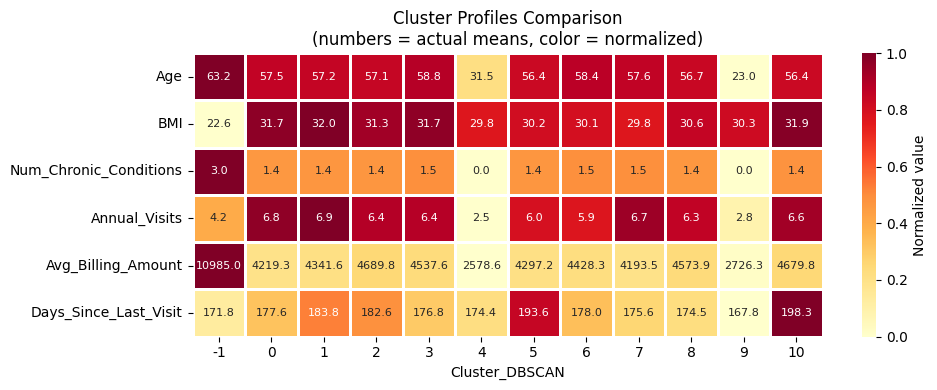
DBSCAN Results#
Why DBSCAN struggles here: High-dimensional patient data (23 features after encoding) tends to have roughly uniform density across the feature space. There are no sharp density drop-offs between patient groups. DBSCAN excels when clusters visibly differ in local density (e.g., spatial or sensor data). Adjusting eps changes the result, but no clear multi-cluster structure emerges.
Key observations:
DBSCAN finds very few meaningful clusters, decreasing
epstends to create lots of small clustersThe noise percentage is sensitive to
eps: increasingepsreduces noise but merges clustersThis behavior is typical for dense, uniformly distributed tabular health data
Step 4.3: MeanShift#
MeanShift iteratively shifts each data point towards the local density maximum (mode) by averaging over all points within a kernel window. Points that converge to the same mode form a cluster.
Key properties:
No need to specify K, the number of clusters is determined automatically by data density
All points are assigned to a cluster (no explicit noise concept)
The
bandwidthparameter controls the kernel size:Larger bandwidth: fewer, coarser clusters
Smaller bandwidth: more, finer clusters
Estimate Bandwidth#
The estimate_bandwidth function uses a nearest-neighbor heuristic to suggest a reasonable bandwidth. The quantile parameter (0–1) controls the fraction of pairwise distances used: lower values yield smaller bandwidth and more clusters.
from sklearn.cluster import MeanShift, estimate_bandwidth
# Estimate bandwidth on a random subset for speed
bandwidth = estimate_bandwidth(X_scaled, quantile=0.1, n_samples=1000, random_state=42)
print(f"Estimated bandwidth: {bandwidth:.4f}")
print(f"\nAdjust quantile to control cluster granularity:")
print(f"- Lower quantile -> smaller bandwidth -> more clusters")
print(f"- Higher quantile -> larger bandwidth -> fewer clusters")
Estimated bandwidth: 5.3331
Adjust quantile to control cluster granularity:
- Lower quantile -> smaller bandwidth -> more clusters
- Higher quantile -> larger bandwidth -> fewer clusters
Fit MeanShift#
ms = MeanShift(bandwidth=bandwidth, bin_seeding=True)
ms_labels = ms.fit_predict(X_scaled)
df['Cluster_MeanShift'] = ms_labels
n_clusters_ms = len(set(ms_labels))
print(f"MeanShift (bandwidth={bandwidth:.4f})")
print(f"Clusters found: {n_clusters_ms}")
print(f"\nCluster sizes:")
for label in sorted(set(ms_labels)):
count = (ms_labels == label).sum()
print(f" Cluster {label}: {count} ({count / len(ms_labels) * 100:.1f}%)")
if n_clusters_ms > 1:
sil_ms = silhouette_score(X_scaled, ms_labels)
print(f"\nSilhouette Score: {sil_ms:.4f}")
MeanShift (bandwidth=5.3331)
Clusters found: 2
Cluster sizes:
Cluster 0: 1934 (96.7%)
Cluster 1: 66 (3.3%)
Silhouette Score: 0.2212
Profile MeanShift Clusters#
We inspect the cluster profiles to understand the characteristics of each patient segment - looking at demographics, health metrics, and healthcare utilization.
# Print cluster profile
print_cluster_profile('Cluster_MeanShift')
| Age | BMI | Num_Chronic_Conditions | Annual_Visits | Avg_Billing_Amount | Days_Since_Last_Visit | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| mean | std | mean | std | mean | std | mean | std | mean | std | mean | std | |
| Cluster_MeanShift | ||||||||||||
| 0 | 51.64 | 14.80 | 30.75 | 8.84 | 1.12 | 0.88 | 5.56 | 3.51 | 4043.75 | 2478.82 | 180.51 | 104.75 |
| 1 | 23.03 | 4.06 | 30.33 | 9.02 | 0.00 | 0.00 | 2.85 | 0.98 | 2726.32 | 1471.93 | 167.76 | 102.86 |
# Print statistics for cluster n
print_cluster_statistics('Cluster_MeanShift', 0)
======================================================================
CLUSTER 0 (n=1934, 96.7% of patients)
======================================================================
Age: 51.6 ± 14.8
BMI: 30.8 ± 8.8
Chronic conditions: 1.12 (avg)
Top conditions: None (429), Hypertension (210), Obesity (183)
Annual visits: 5.6
Avg billing: $4,044
Preventive care: 46.8%
Gender: Male: 50%, Female: 50%
Insurance: Medicare: 47%, Private: 28%, Medicaid: 25%
plot_cluster_means_heatmap('Cluster_MeanShift')
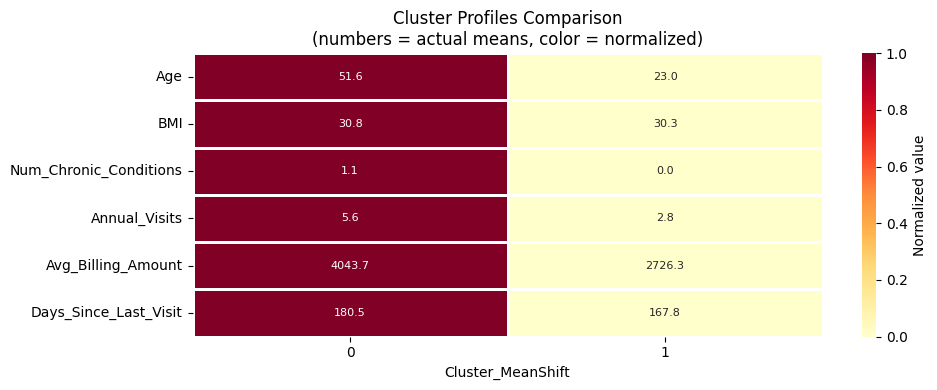
MeanShift Results#
MeanShift often yields a small number of clusters on this dataset because the patient data occupies a relatively compact, uniform region in high-dimensional space.
The clusters found correspond to the dominant density modes.
Observations:
The number of clusters is determined automatically based on data density
Unlike DBSCAN, all patients are assigned to a cluster (no noise)
The bandwidth parameter is the main tuning lever;
estimate_bandwidthprovides a data-driven starting point
Step 5: Comparison of Clustering Algorithms#
We now compare all three algorithms side-by-side using two dimensionality reduction techniques:
Step 5.1: PCA - linear projection, preserves global variance structure, fast
Step 5.2: t-SNE - non-linear projection, preserves local neighborhood structure, slower
Both projections are computed on the same scaled data X_scaled and colored by each algorithm’s cluster labels.
PCA Projection#
from sklearn.decomposition import PCA
# Compute 2D PCA projection
pca = PCA(n_components=2, random_state=42)
X_pca = pca.fit_transform(X_scaled)
print(f"PCA explained variance: {pca.explained_variance_ratio_.sum()*100:.1f}% total "
f"(PC1: {pca.explained_variance_ratio_[0]*100:.1f}%, "
f"PC2: {pca.explained_variance_ratio_[1]*100:.1f}%)")
PCA explained variance: 25.0% total (PC1: 16.1%, PC2: 9.0%)
# Helper: scatter plot of cluster labels on 2D coordinates
def plot_clusters_2d(ax, X_2d, labels, title, xlabel='PC_1', ylabel='PC_2'):
palette = plt.cm.tab10.colors
for i, label in enumerate(sorted(set(labels))):
mask = np.array(labels) == label
color = 'lightgrey' if label == -1 else palette[i % len(palette)]
lname = 'Noise' if label == -1 else f'Cluster {label}'
mk = 'x' if label == -1 else 'o'
ax.scatter(X_2d[mask, 0], X_2d[mask, 1],
c=[color], label=lname, alpha=0.5, s=20,
marker=mk, linewidth=0.2)
ax.set_xlabel(xlabel, fontsize=10)
ax.set_ylabel(ylabel, fontsize=10)
ax.set_title(title, fontsize=12, fontweight='bold')
ax.legend(fontsize=8, loc='best', markerscale=1.5)
fig, axes = plt.subplots(1, 3, figsize=(20, 6))
# KMeans
plot_clusters_2d(ax=axes[0], X_2d=X_pca, labels=kmeans_labels,
title=f'KMeans (K={K})')
# DBSCAN
n_db = len(set(dbscan_labels)) - (1 if -1 in dbscan_labels else 0)
plot_clusters_2d(ax=axes[1], X_2d=X_pca, labels=dbscan_labels,
title=f'DBSCAN (eps={eps_dbscan}, min_s={min_samples_dbscan})\n'
f'{n_db} cluster(s), {n_noise} noise points')
# MeanShift
n_ms = len(set(ms_labels))
plot_clusters_2d(ax=axes[2], X_2d=X_pca, labels=ms_labels,
title=f'MeanShift (bw={bandwidth:.2f})\n{n_ms} cluster(s)')
plt.suptitle('PCA Projection', fontsize=15, fontweight='bold')
plt.tight_layout()
plt.show()
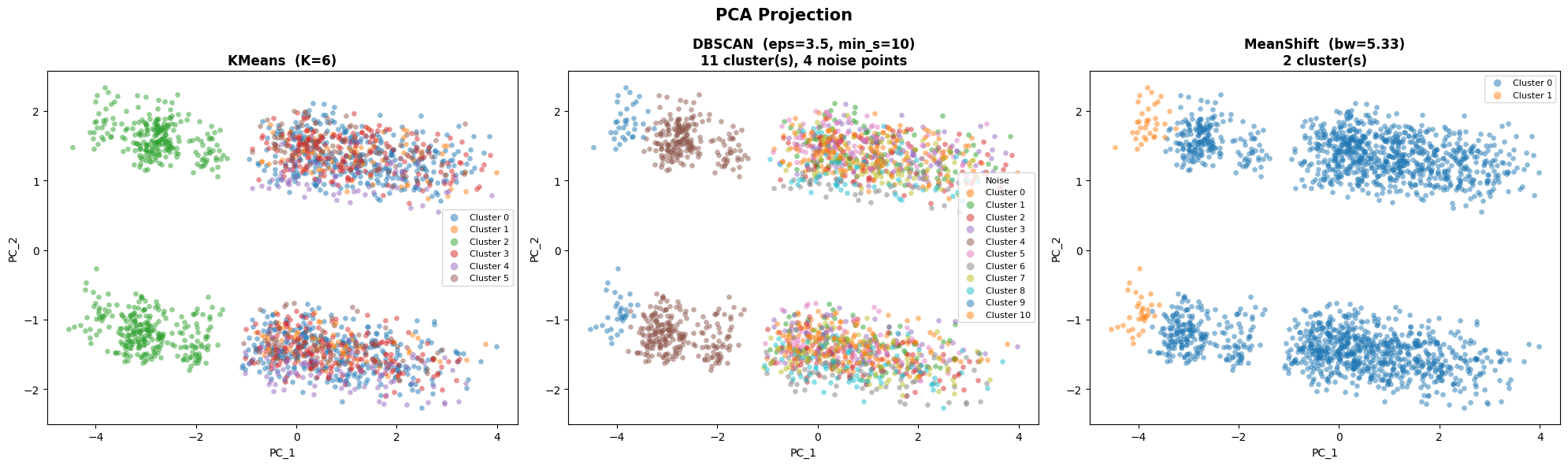
t-SNE Projection#
t-SNE (t-distributed Stochastic Neighbor Embedding) is a non-linear technique that preserves local neighborhood structure, revealing fine-grained cluster topology that PCA may miss.
Key properties:
Non-linear: better at revealing local cluster boundaries
Stochastic: results depend on the random seed and
perplexity(~5–50)Not suitable for projecting new points — purely a visualization tool
Interpretation: nearby points in t-SNE space are locally similar, but distances between clusters are not directly meaningful
from sklearn.manifold import TSNE
print("Computing t-SNE (this may take a moment)...")
tsne = TSNE(n_components=2, random_state=42, perplexity=30)
X_tsne = tsne.fit_transform(X_scaled)
print("Done.")
Computing t-SNE (this may take a moment)...
Done.
fig, axes = plt.subplots(1, 3, figsize=(20, 6))
n_db = len(set(dbscan_labels)) - (1 if -1 in dbscan_labels else 0)
n_ms = len(set(ms_labels))
# KMeans
plot_clusters_2d(ax=axes[0], X_2d=X_tsne, labels=kmeans_labels,
title=f'KMeans (K={K})', xlabel='t-SNE 1', ylabel='t-SNE 2')
# DBSCAN
plot_clusters_2d(ax=axes[1], X_2d=X_tsne, labels=dbscan_labels,
title=f'DBSCAN (eps={eps_dbscan}, min_s={min_samples_dbscan})\n'
f'{n_db} cluster(s), {n_noise} noise points',
xlabel='t-SNE 1', ylabel='t-SNE 2')
plot_clusters_2d(ax=axes[2], X_2d=X_tsne, labels=ms_labels,
title=f'MeanShift (bw={bandwidth:.2f})\n{n_ms} cluster(s)',
xlabel='t-SNE 1', ylabel='t-SNE 2')
plt.suptitle('t-SNE Projection', fontsize=15, fontweight='bold')
plt.tight_layout()
plt.show()
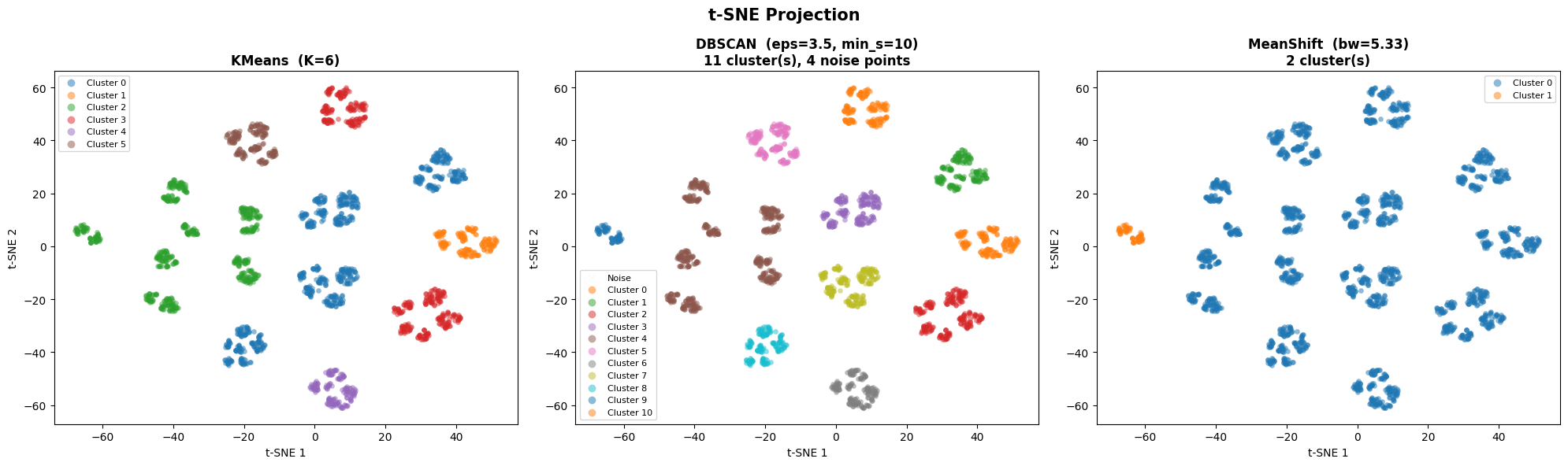
from sklearn.metrics import davies_bouldin_score
results = {}
# KMeans
results['KMeans'] = {
'n_clusters': K,
'n_noise': 0,
'silhouette': round(silhouette_score(X_scaled, kmeans_labels), 4),
'davies_bouldin': round(davies_bouldin_score(X_scaled, kmeans_labels), 4),
'inertia': round(kmeans_final.inertia_, 1),
}
# DBSCAN
mask_nn = dbscan_labels != -1
if n_clusters_dbscan > 1 and mask_nn.sum() > 1:
sil_db = round(silhouette_score(X_scaled[mask_nn], dbscan_labels[mask_nn]), 4)
dbi_db = round(davies_bouldin_score(X_scaled[mask_nn], dbscan_labels[mask_nn]), 4)
else:
sil_db = float('nan')
dbi_db = float('nan')
results['DBSCAN'] = {
'n_clusters': n_clusters_dbscan,
'n_noise': n_noise,
'silhouette': sil_db,
'davies_bouldin': dbi_db,
'inertia': float('nan'),
}
# MeanShift
results['MeanShift'] = {
'n_clusters': n_clusters_ms,
'n_noise': 0,
'silhouette': round(silhouette_score(X_scaled, ms_labels), 4) if n_clusters_ms > 1 else float('nan'),
'davies_bouldin': round(davies_bouldin_score(X_scaled, ms_labels), 4) if n_clusters_ms > 1 else float('nan'),
'inertia': float('nan'),
}
df_metrics = pd.DataFrame(results).T
df_metrics.index.name = 'Algorithm'
print("Quantitative Metric Comparison\n")
print(df_metrics.to_string())
print("\nMetric interpretation:")
print(" Silhouette: higher is better (range: −1 to 1)")
print(" Davies-Bouldin: lower is better (range: 0 to ∞)")
print(" Inertia: lower is better (KMeans only)")
Quantitative Metric Comparison
n_clusters n_noise silhouette davies_bouldin inertia
Algorithm
KMeans 6.0 0.0 0.2034 2.2101 31437.0
DBSCAN 11.0 4.0 0.3233 1.3009 NaN
MeanShift 2.0 0.0 0.2212 1.0641 NaN
Metric interpretation:
Silhouette: higher is better (range: −1 to 1)
Davies-Bouldin: lower is better (range: 0 to ∞)
Inertia: lower is better (KMeans only)
Step 6: Wrap-Up and Summary#
Algorithm Comparison Summary#
KMeans |
DBSCAN |
MeanShift |
|
|---|---|---|---|
Clusters found |
6 (specified) |
Medium (auto) |
Very Few (auto) |
Noise handling |
None, all points assigned |
Labels outliers −1 |
None, all points assigned |
Shape flexibility |
Spherical |
Arbitrary |
Arbitrary |
Main tuning lever |
K |
|
|
Best suited when |
K is known, compact clusters |
Data has clear density contrasts |
Data has clear density modes |
Why KMeans Works Best Here#
Patient data in high-dimensional, mixed-feature space tends to be uniformly dense.
There are no sharp density drop-offs between patient groups. DBSCAN and MeanShift struggle in this setting because:
DBSCAN requires density contrasts that simply don’t exist in this tabular data
MeanShift’s automatic bandwidth estimation yields very few, coarse clusters
KMeans provides stable, interpretable segments despite its spherical-cluster assumption.
Limitations#
KMeans on one-hot data: Euclidean distance on binary features is pragmatic but imperfect; K-Prototypes handles mixed types more naturally
K selection is ambiguous: different K values yield different but potentially valid segmentations — always validate with domain experts
Geographic features excluded: including location could reveal regional patterns
Synthetic data: patterns may be clearer than in real clinical data
Next Steps#
Validate clusters with clinical domain experts
Try K-Prototypes (native mixed-type support) or hierarchical clustering with Gower distance
Apply UMAP for non-linear dimensionality reduction before clustering
Use cluster labels as features for downstream prediction tasks
